CryoARM300 data collection SPA (Alternative)
Setup of SPA data collection with SerialEM Alternative
- Post bake out
- Clipping cartridges
- Taking over the microscope
- Sample insertion/removal
- Prepare gain reference
- CL Stigmator correction
- Pivot point alignment
- Refine Beam shift calibration
- Calculate coma vs IS matrix
- Setting up grid atlas
- Perform test shots
- Square montaging
- Define multi hole pattern
- Finding and combining holes
- Coma free alignment
- View vs Record alignment
- Starting up data collection
- Unclipping cartridges
- Bake out
- Common problems
- Change Liquid nitrogen tank
- Gatan Camera crashes and recovery
Post bake out
This should be only done by experience facility staff and not by user
Cooling and conditioning after bake out
- Start refilling of both Liquid nitrogen tanks by clicking “Auto” in the “LN2 Auto Refill Maintenance window” for “transfer tank” and “stage tank”.
- In TEM center on the microscope pictogram click on “HT”.
- In the operation for “Conditioning Rod” click on “Condition”.
- Once the rod is in position (pink progress bar), click in the line “HT” the button “ON”
- Wait until procedure is finished (~55 minutes).
- If you realize that one of the nitrogen tanks gets refilled start manually also the refill of the second tank by clicking “Refill” in the “LN2 Auto Refill Maintenance” window.
- Once finished HT will be back to 0 kV.
- Switch the “conditioning Rod” back to “Operate”.
- Click on “HT” the button “ON”
- Wait until procedure is finished (~25 minutes).
- close HT window
- In the microscope pictogram hover over “Emission” and click on “Flashing” the button “High”.
- When done click in the same window on “Auto Emission” “ON”.
- Emission will ramp up to 10 uA.
Gun Alignment
DO NOT DO THIS EVEN AS A USER OR SUPERUSER!!!
- Open the beam valve
- Switch off Low Dose mode in SerialEM (if it is on).
- Go to Spot size 1.
- Set Beam Align 1 to Default.
- Go to 10k or 20k Magnification
- Set exposure time on large phosphor screen to 0.001s (minimum)
- Remove CL aperture.
- Condense beam.
- Center beam with GunAlign 1. Focus on the bright spot as center.
- Switch on Anode wobbler.
- Make beam concentric between the changes using GunAlign2. (Spreading in all direction equivalently).
- Repeat GunAlign1 and Anode wobbler until you do not have to reposition GunAlign1 anymore.
- Go to the Spot size you will be working with (i.e. 6)
- Go to 60k Magnification.
- Insert CL aperture.
- Set exposure time on large phosphor screen to 0.02 s.
- Switch on Anode wobbler.
- Adjust GunAlign2 so that the beam is concentric between the changes.
- Control the concentricity with CL aperture removed (Above procedure only works well if CL aperture is centered). Adjust GunAlign2 if necessary.
Clipping cartridges
Note: All images are shown without liquid nitrogen in the workstation for easier visibility.
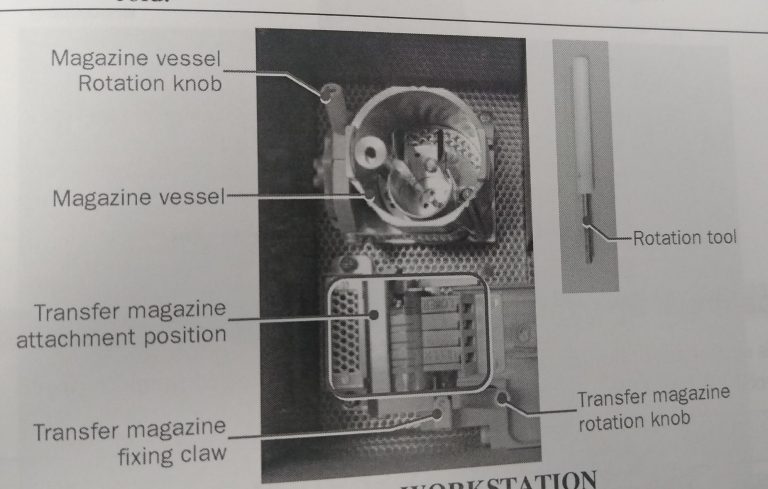




- You can find an empty magazine in the vessel wrapped in aluminium foil in the oven. Never touch it bare handed.
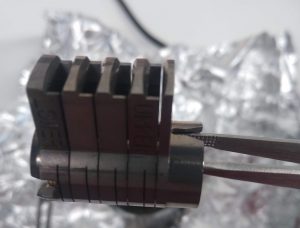


- You can pick up the magazine by putting a tweezer through the two front holes and grab it. (Figure 5 and 6)


- Now grab the magazine with the white pick up tool as shown in Figure 7 and 8


- Put the magazine into the proper position in the workstation. If done well, the magazine is sliding smoothly along the rail (Figures 9 and 10).
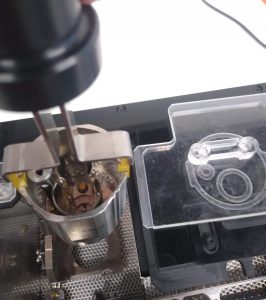

- Grab the cup tightly with the carrying tool and insert it into the workstation. Use the pictogram as a guide to know the proper rotation. It should sit tight (Figure 11).
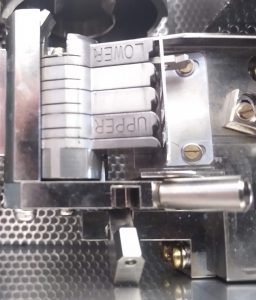
- Cool down the workstation with liquid nitrogen. Once cold, make sure that the liquid nitrogen level is not higher than the specimen attachment area (Figure 1).

- Use the white rotation tool (Figure 1) to turn the cup 90 degrees with the opening towards the magazine (Figure 12).



- Use the white rotation tool to turn the magazine ~90 degrees with the opening towards the assembly site (Figure 13). Once in position (Figure 14), engage the safety pin (Figure 15)

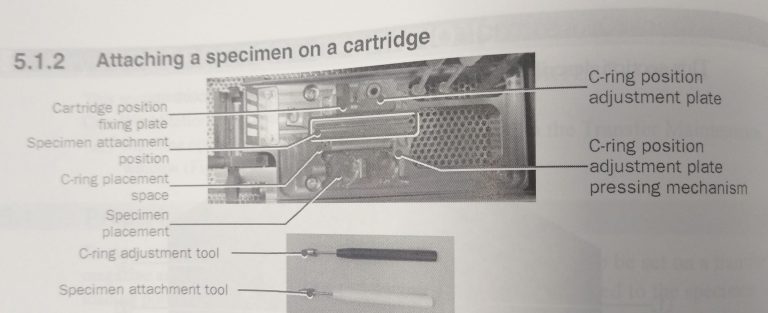
- Place the empty cartridge in the cartridge placement space (Figure 16).
- Place your grid box in the specimen placement area (Figure 16).
- Place the C-ring in the C-ring placement space (Figure 16).

- Ensure that the tip of the specimen attachment tool is properly spread by inserting the rotation tool at the end (Figure 17).

- Put a tweezer, the black C-ring adjustment tool and the white specimen attachment tool into the workstation and make sure they are cooled down before use (Figure 18).

- Put a cartridge with the screw facing you and the square toward the left on the rail of the specimen attachment area (Figure 19). If properly positioned, the cartridge can slide easily sideward.
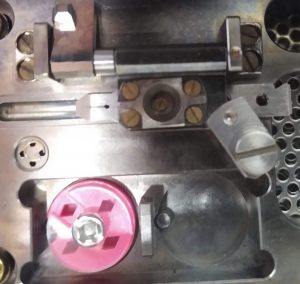

- Align the screw of the cartridge, with the hole of the C-ring adjustment plate and put down the plate. Lock it with the rotation lock (Figure 20). Note: The rotation lock here is not essential but helpful

- Insert one C-ring into the C-ring adjustment plate (Figure 21). Note: The C-ring does not need to be completely flat in position, but not totally tilted either.

- Use the black C-ring adjustment tool to clamp the C-ring in the C-ring adjustment plate by pushing the tool straight down (Figure 22).

- Lift up the C-ring attachment plate and control that the C-ring is properly sitting in the plate (Figure 23). Important: It needs to be straight, otherwise clipping will fail.
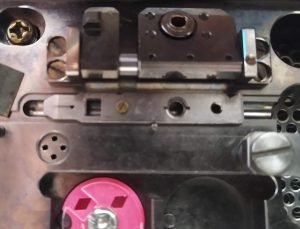

- Move the cartridge further left on the attachment position, so the open square is aligned with the square lock (Figure 24).
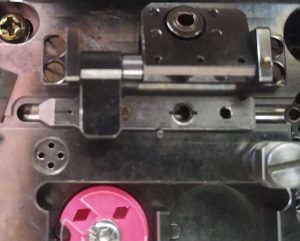

- Lock the cartridge with the square lock (Figure 25). The cartridge should not be able to move sideward anymore.

- Put your cryo grid, top side upwards, onto the cartridge. Make sure its properly positioned in the groove (Figure 26)
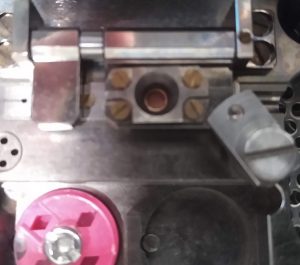

- Put down the C-ring attachment plate with the C-ring, covering the cryo grid. It is vital to use the rotation lock to rigorously lock the attachment in position (Figure 27).

- Use the white specimen attachment tool to clip the C-ring onto the cartridge, securing the cryo grid. Important: Come down straight with the tool, no twisting, no turning (Figure 28).

- Open the rotary lock and carefully lift up the attachment plate. If the clipring is not completely on the cartridge you can put down the attachment plate again and use the white tool again for clipping. Otherwise you can lift up the plate completely. Important: Carefully check that the clipring is properly and straight on the cartridge. (Figure 29)

- Open the square lock from the cartridge (Figure 30).

- Slide the cartridge further left. It will automatically tilt toward you at the position shown in Figure 31.

- Pick up the cartridge with a blunt tweezer and let it glide into the magazine (Figure 32). In the depicted image it is put in position 4.

- Use the straight edge of the white rotation tool to put the cartridge tightly into the magazine. The is a small resistance to overcome (Figure 33).
- Repeat above steps for up to 4 samples.

- Open the lock of the magazine (Figure 34).

- Use the white rotation tool to turn the magazine down (Figure 35).

- Use the flat side of the rotation tool to push the magazine into its vessel (Figure 36).

- Once the magazine is in its vessel use the white rotation tool to lift up the vessel (Figure 37).

- Once the magazine vessel is upright, fill it up with liquid nitrogen, so that the whole magazine is covered. No need to fill it up to the rim of the vessel (Figure 38) Important: Before doing the next steps, make sure the microscope is ready for grid insertion, i.e. it is not making images at the moment.

- Put the cryo cup next to the assembly workstation, have the pictogram on the lid oriented in the same direction as the magazine vessel; i.e. small and large circular structure (Figure 39)

- Pick up the magazine vessel with the large grabbing tool (Figure 40). Important: Ensure a good grid to not drop the vessel.
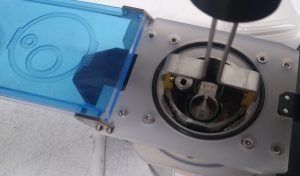

- Position the magazine vessel in the cryo cup (Figure 41). Note the proper orientation with the pictogram.

- Close the lid of the cryo cup and bring it to the microscope for insertion.
- Clean up after finishing and put all the tools back into its box and drawer.
Taking over the microscope
- Stopping the currently running data collection.
- In SerialEM click on “End Navigator” (Dark Green Panel) and wait until the last image in the cycle is recorded.
- Save navigator file by clicking “Navigator –> Save”.
- Save log file by clicking “File –> Save Log”.
- Save settings file by clicking “Settings –> Save”.
- Set Image shift back to 0 by clicking “Reset Image shift” (yellow panel).
- Check when liquid nitrogen refill is happening next. The ideal goal is to either do the liquid nitrogen refill now or after the last coma free alignment (latest in 7h).
- Open the Liquid nitrogen monitor by right click on the microscope scheme in the TEM center and left click on “LN2 auto Refill Maintenance” (Figure 1)
- If it is less than 80% full, click on “refill” for “stage tank” and “transfer tank” (Figure 2)
- Wait until nitrogen filling is finished
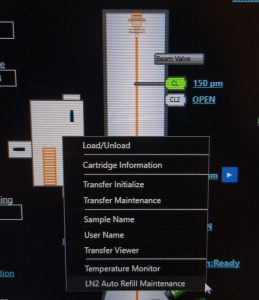

- On the microscope PC, if TEM center is not open, open TEM Center Suite launcher (Figure 3).

- On the camera PC, open the SerialEM executable with your desired imaging conditions (i.e. magnification and CDS/non-CDS mode) (Figure 4). We use mainly the executable 60k_CDS.

- If not yet open, open Digital Micrograph, on the camera PC by clicking on the “GSM3” executable (Figure 5).

- If emission is on (visible in TEM center, on microscope scheme top right, Figure 6): In TEM center hover your mouse over “Emission” and click on “Auto Emission –> Off” (Figure 7). Wait until finished. Afterwards hover your mouse over “Emission” and click “Flashing High” (Figure 7). Once finished click on “Auto Emission –> ON”. Wait until the Emission is ramped up (It will show 10 uA). Continue at point 8. Else follow point 7.
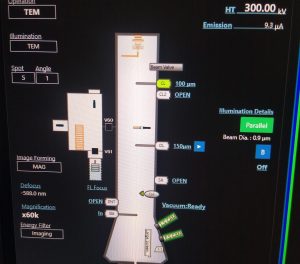
- If emission is off: In TEM center hover your mouse over “Emission” and click on “Flashing High” (Figure 7). Once finished click on “Auto Emission –> ON”. Wait until the Emission is ramped up (It will show 10 uA).
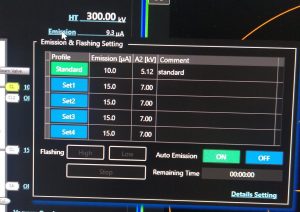
- Degauss the omega filter. Open in the TEM center the window by clicking in the menu on “Maintenance –> Filter Setting” (Figure 8).
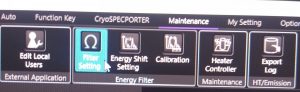
- In the Window Filter Setting click on “Degauss –> Start” (Figure 9). Note: The FL focus value is now offset and the filter will relax in the following hours. During most procedures that follow first, you can remove the Energy filter slit first, and only adjust FL focus once its really needed and probably more stable.

- In SerialEM, make sure you are in “Low Dose mode” and load “Record” mode (pink window, Figure 10).
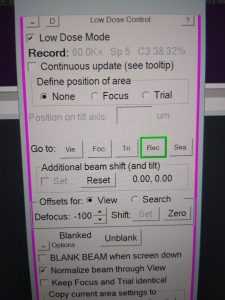
- If not yet open, open the “CryoSpecporter” window by right click on “microscope scheme (TEM center) –> Load/Unload (left click)” (Figure 11).

- Remove the grid from the stage by highlighting the stage sample and clicking on “Keep to storage” (CryoSpecporter window, Figure 12). Wait until the process is finished.
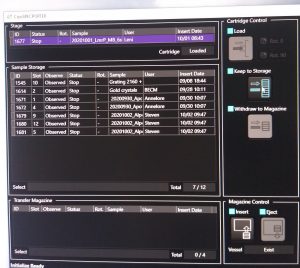
Sample insertion/removal
- Remove previous samples. If the magazine is still present in the microscope, select in the “CryoSpecporter” window up to 4 samples to remove and click on “Withdraw to Magazine” (Figure 1).
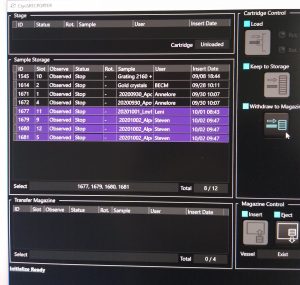
- Once the cartridges are in the transfer magazine click in the “CryoSpecporter” window on “Eject” (Figure 2).
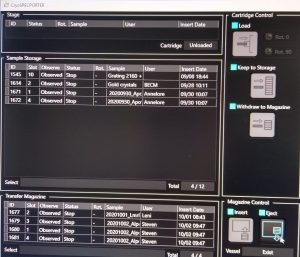
- In the newly opened window click “Yes” (Figure 3) and wait until the cup with the magazine dropped.
- Now you can physically remove the mug from the microscope (Figure 4&5).


- Insert the mug with your new cartridges into the microscope by pushing the mug deep into the system. The lid will slide of automatically (Figure 6).

- On the touch panel on the right hand side touch to wake up the display and select “Insert Magazine” (Figure 7).

- Once the magazine is in the microscope the cartridges will be recognized by the microscope and can be seen in the “Transfer Magazine” part of the “Cryospecporter” window (Figure 8). Then the cartridges will be moved into the sample storage.
- Finally name the cartridges. Right click on the sample –> “cartridge information (left click)” (Figure 9). In the newly open window define “Sample Name” and “User Name” (Figure 10). Note: These names are not used automatically as file name of your images.
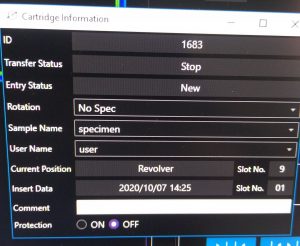
- Repeat the naming for all the cartridges you inserted.
Prepare gain reference
- If there is a grid on the stage, highlight it on the CryoSpecporter window and click "Keep to storage" (Figure 1)
- Open Beam Valve with the “Beam” button (left hand panel).
- In SerialEM make one view image by clicking “View” (dark green panel) to ensure that the camera is in CDS or non-CDS mode (in case you use a different mode than the user before you). Note: Sometimes the camera temperature is still equilibrating (warning appears after attempting to make the image). In this case wait 5 seconds a redo the view image.
- Switch off low dose mode in SerialEM (pink panel).
- Remove the energy filter slit by pushing the “Slit In” button (right hand panel)
- Put down the large phosphor screen (Screen retract, right hand panel).
- If inserted, remove the OL aperture (double click on OL in microscope pictogram)
- Change spot size to 1 using the “Spot size” dial (left hand panel)
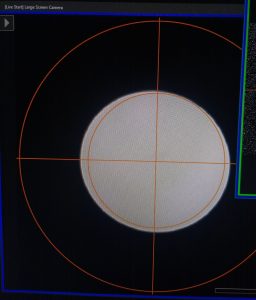
- Center the beam with the “Shift X&Y” dials (both hand panels) and spread the beam until it covers a bit more than the inner circle on the “Large Screen Camera” (Figure 2).
- In Digital Micrograph ensure that the Power User is activated in the menu “Help –> User –> Power User” (Figure 3).

- In Digital Micrograph, make sure that Counting mode is active under "Technique Manager --> TEM Imaging --> Counted" (Figure 4). Hint: If this is not the case the procedure will only do 1 linear gain reference.
- In Digital Micrograph menu select “Camera –> Prepare Gain Reference” (Figure 5).
- Once the Dark Reference is done, lift the Phosphor Screen (“Screen Insert, right hand panel).
- Adjust the beam brightness (left hand panel), so that the measured dose in Digital micrograph corresponds more or less to the one requested (Should be 760). If ready, click on “done” and wait until its done (~8 minutes) (Figure 6).
- Once done, put down the phosphor screen (“Screen retract”, right hand panel).
- Change spot size to 5 (if using CDS mode) or 6 (if using non-CDS mode) using the “spot size” dial (left hand panel).
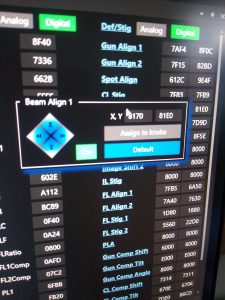
- Roughly center the beam, by hovering in the “Lens/Deflector” window over “Beam Align 1” and click on “Default” (Figure 7).
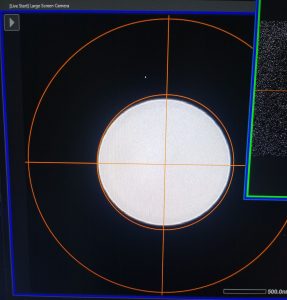
- Center the beam using the “Shift X&Y” dial (both hand panels) and spread the beam (“brightness dial”, left hand panel) until you have the beam at least fully covering the inner circle on the “Large Screen Camera” window (Figure 8).
- In the next window make sure that “expert mode” is not activated and click on “OK” (Figure 9).
- Once the dark reference is done, lift the Phosphor screen (“Screen retract”, right hand panel).
- Adjust the beam brightness (“brightness” dial, left hand panel) until the measured dose correspond pretty well to the requested dose (usually: CDS mode 8.0, non CDS mode 18) (Figure 10).
- Then, confirm the window by clicking “Done”. Wait until the procedure is done (~8 minutes).
- On the next window click “OK” (Figure 11).
- In SerialEM activate “Low Dose Mode” (pink panel, Figure 12) and select record mode (“Rec.”).

- Put down the phosphor screen (Screen retract, Right hand panel).
- Center the beam using the shift X&Y dials (both hand panels).
- In SerialEM click on Setup under the camera& Script panel (dark green) (Figure 13).
- In the new window under the record tab, ensure that "save frames" is not active (no checkbox mark). (Figure 14).
- Click on Acquire to make an image in the empty area. (Figure 14).
- In the SerialEM main window you see a blank image and on the top left corner you can read out the beam intensity (Figure 15)
- Afterwards switch back on "save frames" under the record tab and close the window with "OK"
- Close the beam valve using the “beam” button (left hand panel).
CL Stigmator correction
- Go to record mode (pink panel) (Figure 1).
- Put down the large phosphor screen (screen retract, right hand panel)
- Condense the record beam to a small spot using the brightness dial (left hand panel, counter-clockwise)
- Roughly center the beam using Shift X&Y (both hand panels)
- Insert the focus screen (FScreen retract, right hand panel)
- Roughly center the beam using Shift X&Y (both hand panels)
- If the beam is not circular, assign CL Stigmators to the Def/Stig knobs (CL Stig, left hand panel, will light up). Otherwise you do not need to do this correction.
- Use Def/Stig X&Y knobs to make the beam circular
- Once finished un-assign CL Stigmators by clicking CL Stig (left hand panel)
Pivot point alignment
These pivot points are rather stable and it is recommended to change them only if you get unusually large beam tilt values in the coma vs IS matrix.
Insert grating grid
- Insert alignment grid by highlighting the grating grid (with latex beads) in the CryoSpecporter and click on load. (Figure 1) Note: If there is another grid still on the stage it will be automatically unloaded.
- Once insertion is finished you can open the beam valve. Note: If the beam valve icon is still gray shaded in the pictogram, you still need to wait a few seconds.
Beam tilt pivot points
- Go to standard focus by double clicking “STD Focus” (right hand panel).
- Ensure that you are at eucentric height. Note: Use rough eucentric for a rough targeting. Measure the actual defocus in SerialEM (Menu –> Focus/Tune –> Measure defocus). The log file will display the actual defocus. Use the Z up and Z down buttons (small panel) to adjust the eucentric height and confirm/readjust with “Measure Defocus”. Note: If you measure too fast after stage x,y or z-motion there is still residual drift and this will lead to a mis-estimation of the actual defocus
- Select record beam conditions in SerialEM (pink panel, Rec.).
- Condense the beam to a small point (Brightness dial, left hand panel) and roughly center the beam on the large phosphor screen with “Shift X and Y”(both hand panels) (Figure 2).
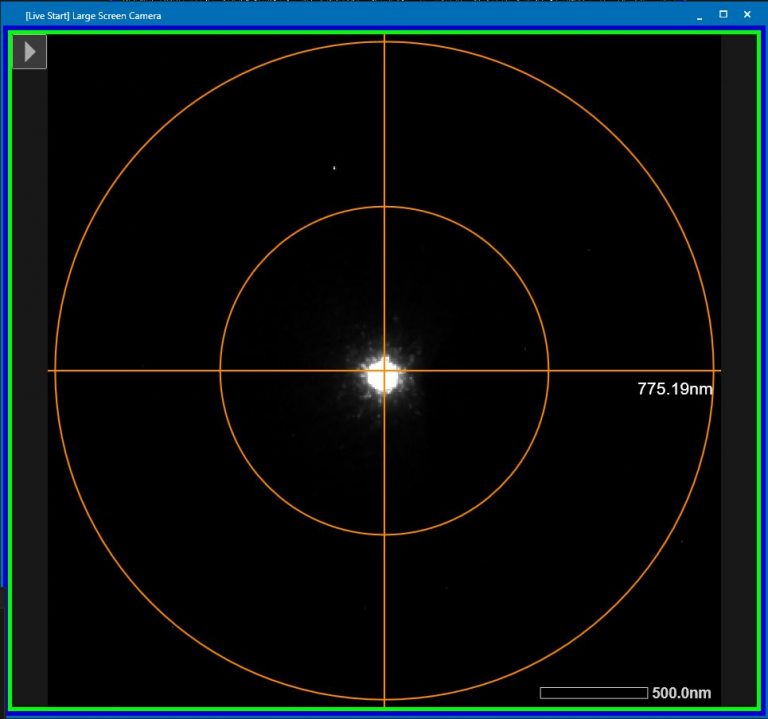
- Insert small phosphor screen “FS retract” right hand panel.
- Condense the beam further into a very small dot (Brightness dial, left hand panel) and center the beam on this Focus screen with “Shift X and Y” (both hand panels) (Figure 3).
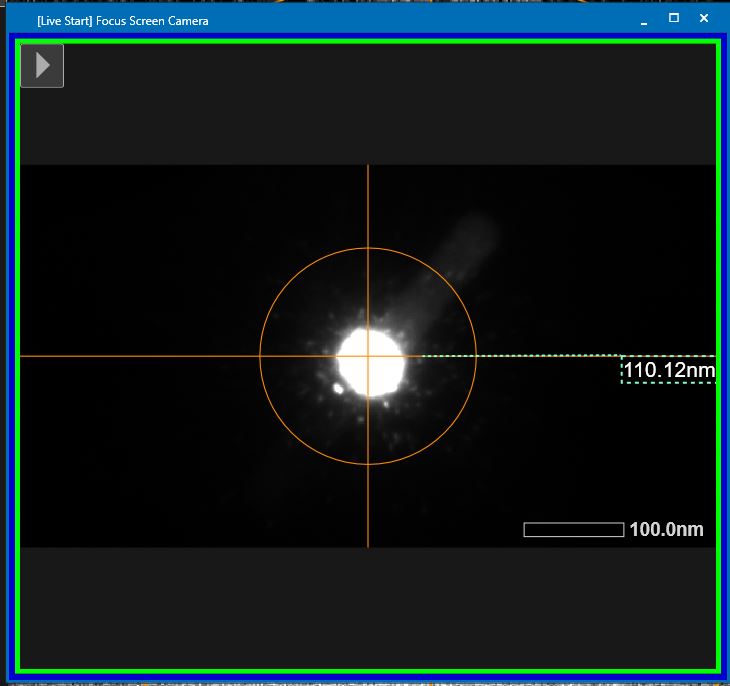
- Open the wobbler window in TEM center through the “Lens/Deflector” window by hovering over “Wobbler” and clicking on “Wobbler window” (Figure 4).
- Select in the new window “Tilt X” (Figure 5). CL Comp tilt will be automatically assigned to the Def/Stig knobs.
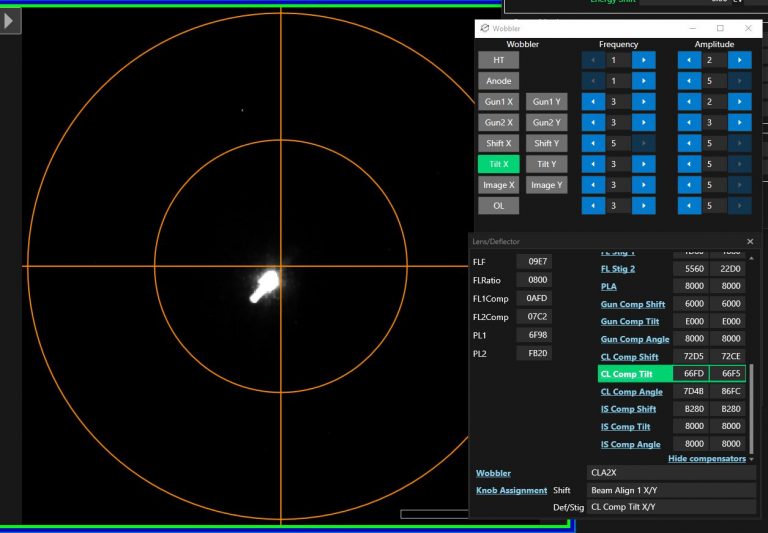
- Use “Def/Stig X” knob (left hand panel) and minimize the displacement of the major beam along the wobbling axis. Note: You can use Beam shift if you will lose the beam from the small Phosphor Screen.
- If there is perpendicular motion to the wobbling axis, CL comp angle needs adjustment. Otherwise continue with point 16.
- Assign CL Comp Angle to the "Def/Stig: knobs via Lens/Deflector window –> CL Comp Angle –> Assign to knobs”.
- Click on one of the “Def/Stig” button to go into coarse mode.
- Use the “Def/Stig X”button to minimize the lateral motion against wobbling axis.
- Click on one of the “Def/Stig” button to go back into fine mode.
- Assign CL Comp Tilt back to the "Def/Stig: knobs via Lens/Deflector window –> CL Comp Tilt –> Assign to knobs”.
- Switch off “Tilt X” and switch on “Tilt Y”.
- Use “Def/Stig Y” knob (left hand panel) and minimize the displacement of the major beam along the wobbling axis. Note: You can use Beam shift if you will lose the beam from the small Phosphor Screen.
- If there is perpendicular motion to the wobbling axis, CL comp angle needs adjustment. Otherwise continue with point 24.
- Assign CL Comp Angle to the "Def/Stig: knobs via Lens/Deflector window –> CL Comp Angle –> Assign to knobs”.
- Click on one of the “Def/Stig” button to go into coarse mode.
- Use the “Def/Stig X”button to minimize the lateral motion against wobbling axis.
- Click on one of the “Def/Stig” button to go back into fine mode.
- Assign CL Comp Tilt back to the "Def/Stig: knobs via Lens/Deflector window –> CL Comp Tilt –> Assign to knobs”.
- Switch off “Tilt Y”.
Beam tilt
As above you still need to be at standard focus and eucentric height.
Ensure that there is not residual drift from sample insertion or liquid nitrogen refilling.
- Remove the Small Focus Screen (“FS insert”, right hand panel)
- Put the beam in parallel mode by loading record mode from SerialEM (“Rec.”, pink panel)
- Center the image with the phosphor screen using the stage on one of the latex beads, center the beam (Figure 6).

- Insert the small focus screen “FScreen retract”, (right hand panel).
- Center the latex bead on the screen using the trackball.
- Paint a circle around the position of the latex bead (TEM Center menu –> Tools –> Ellipse)
- Change the defocus to between -10 to -15 um (“Image/diff focus button”, right hand panel). If there is significant beam tilt you will see that the image starts moving.
- Switch on “Bright tilt” (Left hand panel). This will assign BeamAlign2 to the Def/Stig knobs.
- Center the latex bead again on its original position using “Def/Stig X&Y” buttons (both hand panels, use coarse mode).
- Hit standard focus (Std focus, right hand panel).
- Repeat steps 31-34 until the bead does not change its position during defocusing.
- Switch off “Bright tilt” (Left hand panel).
Beam shift pivot points
From before you should still be at standard focus and eucentric height. You should also still have a latex bead in the center of the focus screen
- Remove condenser lens aperture (CL button, microscope pictogram, TEM center).
- Set defocus to -40 um (“Image/Diff focus” button, right hand panel).
- In the Lens/Deflector window hover over “CL Comp Shift” and push “assign to knobs”.
- In SerialEM edit the script Shift X,Y (menu –> Script –> Edit –> Edit 6 (or similar)) and put amp_X to 40 and amp_Y to 0, then run script Shift X,Y (Figure 7). You may also want to change "loop NUMBER icount" to between 10-20 dpending how fast you can adjust the pivot point.
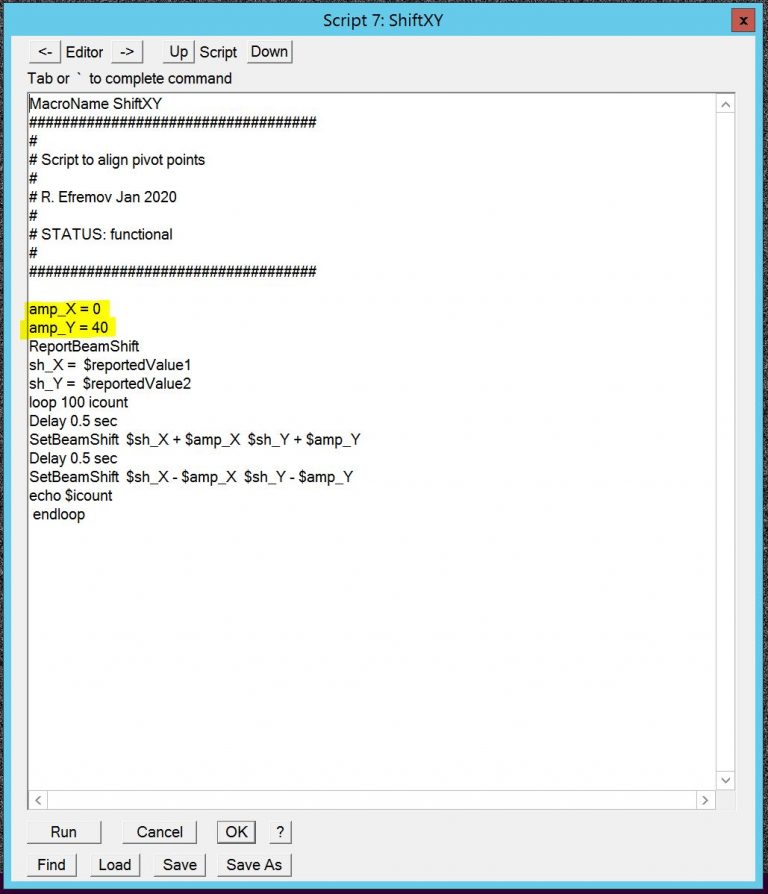
- Minimize the movement of the latex bead along the beam motion on the focusing screen using “Def/Stig X”button. Note: There may be image motion perpendicular to the beam motion. Do not try to correct for this!
- Wait until the script stops automatically (length depends on icount number)
- If you stopped the script accidentally, roughly center the beam via Set BEAM ALIGN 1 to default by hovering in the Lens/Deflector window over Beam Align 1 and hit “Default”.
- Edit script Shift X,Y and put amp_X to 0 and amp_Y to 40, then run script Shift X,Y (Figure 7).
- Minimize the movement of the latex bead along the beam motion on the focusing screen using “Def/Stig Y”button. Note: There may be image motion perpendicular to the beam motion. Do not try to correct for this!
- Wait until the script stops automatically (length depends on icount number)
- If you stopped the script accidentally, roughly center the beam via Set BEAM ALIGN 1 to default by hovering in the Lens/Deflector window over Beam Align 1 and hit “Default”.
- Un-assign “CL Comp Shift” from the knobs (via Lens/Deflector window –> CL Comp Shift –> Assign to knobs”). Button will not be green anymore.
- Insert condenser lens aperture (CL button, microscope pictogram, TEM center).
- Push STANDARD FOCUS (“STD focus”, right hand panel).
- Center the beam with “Shift X&Y” button (both hand panels).
Refine Beam shift calibration
This normally only needs to be done if you changed the pivot points.
- Ensure that there are at least 6 um of grid around your current location, by:
- Select View mode in Serial EM ("Vie., pink panel) (Figure 1).
- Put down the large Phosphor screen ("screen insert", right hand panel).
- Remove the CL aperture (double click on CL in microscope pictogram, TEM center) (Figure 2).
- Move the stage with the tracking ball to a suitable location (Figure 3).
- Re-insert the CL aperture (double click on CL in microscope pictogram, TEM center) (Figure 2).
- In SerialEM go to trial mode ("Tri", pink panel) (Figure 4).
- Go to standard focus by double clicking “STD Focus” (right hand panel).
- Put down the large phosphor screen by clicking "Screen retract" (right hand panel).
- Center the beam with “Shift X&Y” button on the large phosphor screen (both hand panels) (Figure 5).
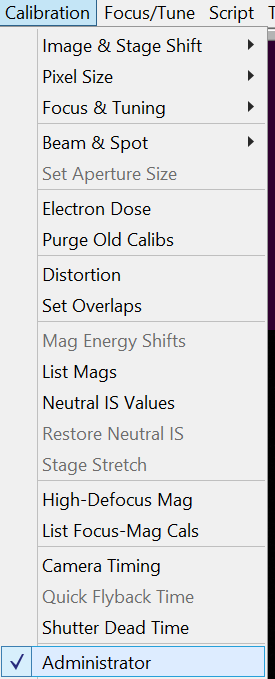
- In SerialEM select in the Menu bar Calibration --> Administrator (Figure 6).
- In SerialEM select in the Menu bar Calibration --> Beam & Spot --> Refine Beam shift (Figure 7).

- Confirm window with "Use Centroid" (Figure 8).
- In the new window, ensure that the selection screen asks for 6 um image shift. Confirm with "OK" (Figure 9).
- The program will first apply a small beam shift in 4 directions and calculate a matrix from the observed beam shift. It will the do the same in 2 rounds with the large image shift. Note: Make sure that the images in all four directions are bright and you are not blocked in calibration by a grid bar.
- The log file will print out the refined matrix numbers. Note: If this changed dramatically from the staring matrix you probably have done something wrong while setting the pivot points (Figure 10).
- Save the calibration in SerialEM by clicking in the Menu bar Calibration --> Save Calibrations (Figure 11).
- Get out of Administrator mode by unselecting in the Menu bar Calibration --> Administrator (Figure 12).
Calculate coma vs IS matrix
It is recommended to do this matrix calculation before deciding to adjust the pivot points. The pivot points are mostly stable. You should run this procedure before each new data collection to ensure that you have a good coma and astigmatism correction during multi-hole recording. The way shown here is only useful if you collect with SerialEM internal scripts only (i.e. this alternative protocol)
- If not yet done, insert alignment grid by highlighting the grating grid (with latex beads) in the CryoSpecporter and click on load. (Figure 1) Note: If there is another grid still on the stage it will be automatically unloaded.
- Ensure that there are at least 6 um of grid around your current location, by first selecting View mode in Serial EM ("Vie., pink panel) (Figure 2).
- Put down the large Phosphor screen ("screen insert", right hand panel).
- Remove the CL aperture (double click on CL in microscope pictogram, TEM center) (Figure 3).
- Move the stage with the tracking ball to a suitable location (Figure 4).
- Re-insert the CL aperture (double click on CL in microscope pictogram, TEM center) (Figure 3).
- Select record beam conditions in SerialEM (pink panel, Rec.) (Figure 5).
- Go to standard focus by double clicking “STD Focus” (right hand panel).
- Ensure that you are at eucentric height. Note: Use rough eucentric for a rough targeting. Measure the actual defocus in SerialEM (Menu –> Focus/Tune –> Measure defocus). The log file will display the actual defocus. Use the Z up and Z down buttons (small panel) to adjust the eucentric height and confirm/readjust with “Measure Defocus”. Note: If you measure too fast after stage x,y or z-motion there is still residual drift and this will lead to a mis-estimation of the actual defocus
- Put down the large phosphor screen by clicking "Screen retract" (right hand panel).
- Center the beam with “Shift X&Y” button on the large phosphor screen (both hand panels) (Figure 6).
- Ensure that there is no residual drift at this moment. Wait until good.
- Define a target defocus by clicking in the SerialEM Menu bar on Focus/Tune --> Set Target (Figure 7).
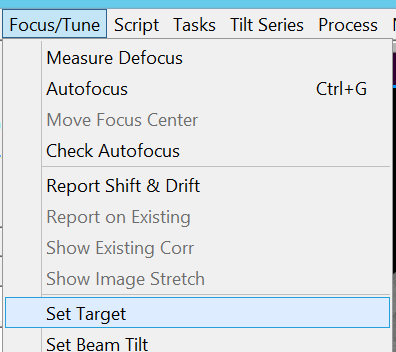
- Set the target to -1.3 um defocus (Figure 8).
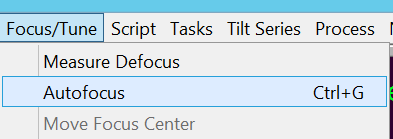
- In the Menu bar go to Focus/Tune --> Autofocus to do auto-focusing (Figure 9). Hint: If the focus position ends up in the hole and focusing fails, you may want to temporarily set "Define position of area -- Focus" to 0 um offset. Undo this before starting the data collection.
- Do coma free alignment by clicking in the Menu bar on "Focus/Tune --> Coma-free alignment by CTF" (Figure 10). Important: Check in the FFT image that the fit (yellow dashed lines) corresponds to measured location of Thon rings.
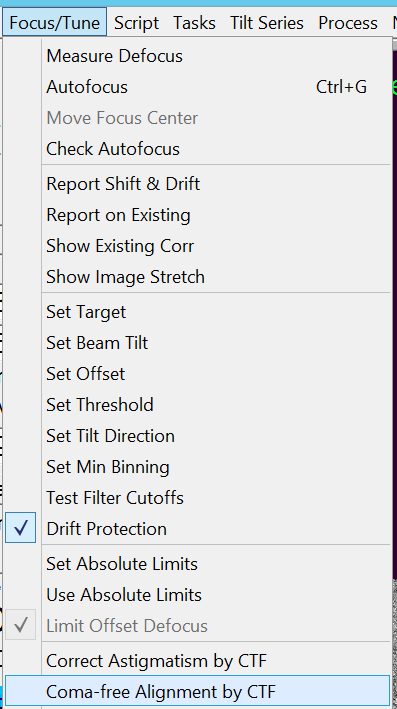
- The log file reports the amount of beam tilt correction (Figure 11). Hint: High score and low fit to values indicate a good run.
- Do astigmatism correction by clicking in the Menu bar on "Focus/Tune --> Coma astigmatism by CTF" (Figure 12).
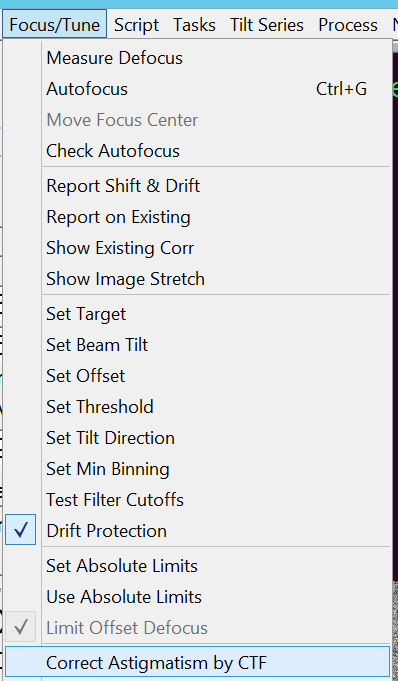
- The log file reports the amount of OL stigmator correction (Figure 13). Hint: High score and low fit to values indicate a good run.
- Run the Coma vs IS calibration by clicking in the Menu Bar "Focus/Tune --> Calibrate Coma vs Image Shift". Important: For each tilt direction control the amount of beam tilt needed. It should be relatively symmetrical and less than 2 mrad. If not: check pivot points. If pivot point are for sure still good, check gun alignment (Gun alignment should be done by staff only!!!).
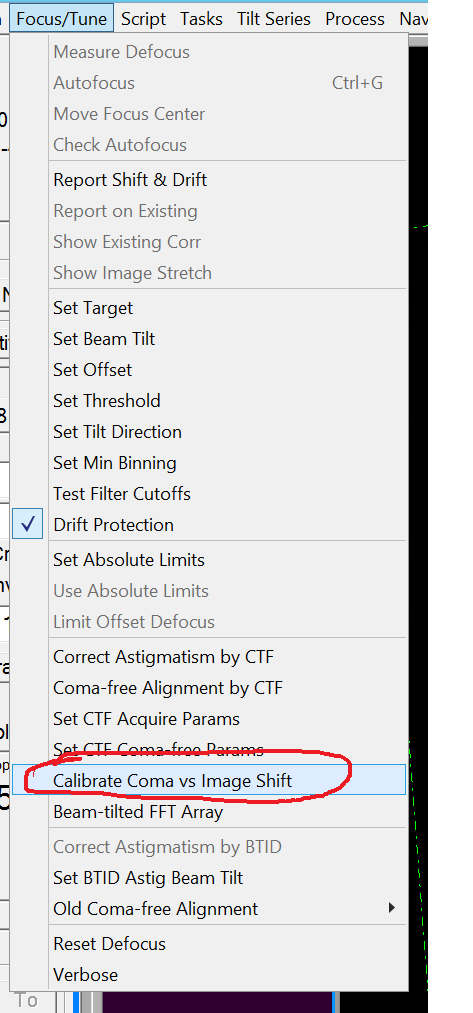
- At the end of the procedure the log file displays values for BT vs IS and STIG vs IS (Figure 15).
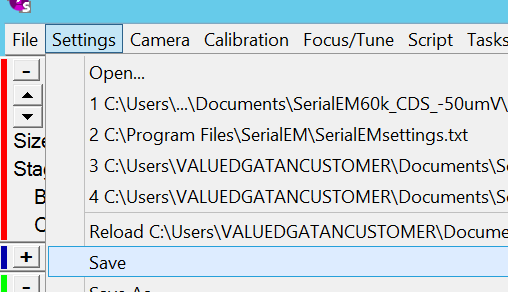
- Save the calibration by clicking in the Menu Bar "Settings --> Save" (Figure 16).
Setting up grid atlas
Note: Following procedure assumes that you are still in Low Dose Mode (pink panel)

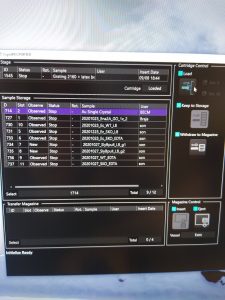
- Insert the grid you want to measure onto the stage by selecting it in the Cryospecporter window and clicking on load (Figure 1). Note: It will automatically unload any sample currently on the stage.
- Open the beam valve (beam button, left hand panel). Note: If the beam valve icon is still gray shaded in the pictogram, you still need to wait a few seconds.
- In the pink SerialEM panel select “Vie.” to load the view microscope parameters (Figure 2).

- Put down the large phosphor screen (right hand panel, screen retract)
- Move the stage with the trackball until you see a square in the center of the field of view (Figure 3). Note: Skip this for now if you can’t find a square close by and do it after the atlas is collected. (Update Z of map)

- Move the stage to eucentric height using serialEM, by clicking in the menu on “Task –> Eucentricity --> Rough Eucentricity” (Figure 4).
- Open a new navigator window via “Menu –> Navigator –> Open” (Figure 5).
- Open a new log file via “Menu –> File –> Open Log” (Figure 6).
- To load some basic parameters, load an old atlas via the Menu bar File --> Open Old (Figure 7). Note: Step 9-12 This only has to be done once if you do multiple grid montages in a row without setting up a square montage
- Find under CaptureData(X:)\Serial_EM_raw_data\Standard_atlas the file atlas_bin2_DATE.mrc (Figure 8&9). Open file by clicking "open".
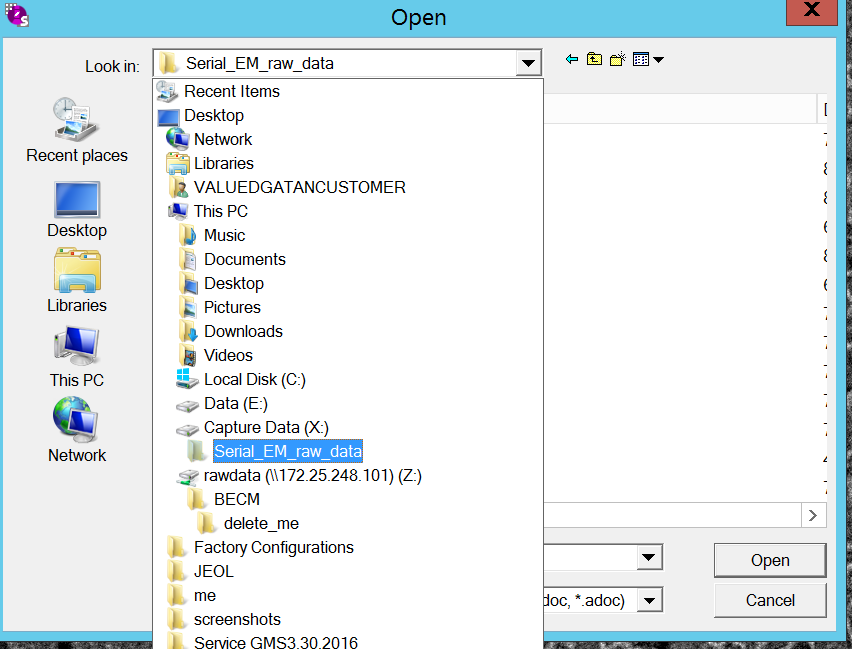
- In the new conformation window select "Yes" (Figure 10).
- In the next window ensure that "Move stage instead ..." and "Use continuous mode ... 0.5" are checked (Figure 11). Confirm with "Ok'
- Click in the scripts panel on “MakeGridAtlas” to start the atlas collection procedure (Figure 12).
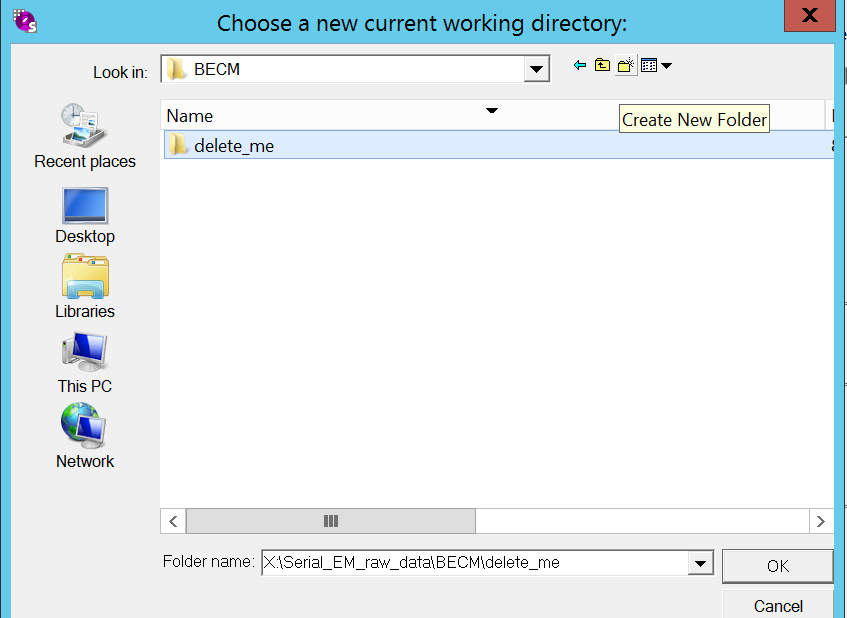
- Create the a new folder in your personal working directory under “X:\Serial_EM_raw_data\myGroup\myName”, give it a good name and click “OK”(Figure 8). Important: Ensure you are under the X drive. NEVER Z!!!
- Once the atlas is collected you may hear a beep. You can ignore this. A new window will appear to save the navigator file. Ensure you are in your working directory (Still under X:) and save the file as “navigator”. Click “Save” (Figure 14).
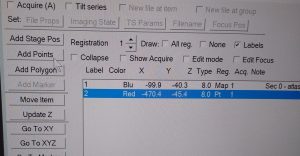
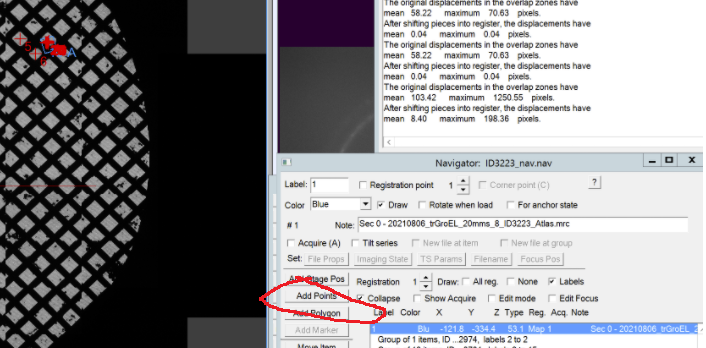
- In the navigator window a Blue labelled line will appear with the name “Sec 0- Atlas.mrc”. Double click on this line to open the map in a higher resolution (Figure 15). Note: You may use the option "align pieces in overview" in the dark brown panel. You need to reopen the map after activating this option
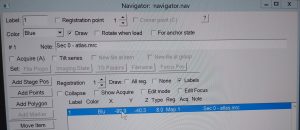
- In the navigator window select “Add points” and select in the map an easily identifiable feature. Left click on that feature add the point in the navigator window. Finally click “Stop Adding” in the Navigator window (Figure 16).
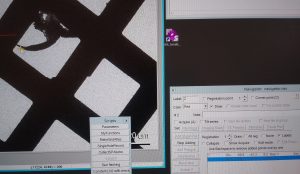
- Make sure that the newly added point is highlighted in the Navigator window and select “Go to XYZ” (Figure 17).
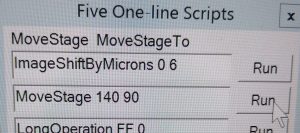
- In the One-line Scripts window “Run” the script “MoveStage 15 90” (Figure 18). Note: The recommended offset changes over longer time and may be adjusted if necessary.
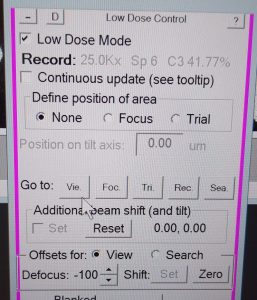
- In the pink SerialEM panel select “Go to”: “Vie.” (This is view mode) (Figure 19).
- Remove the CL aperture by double clicking on “CL” in the microscope pictogram in “TEM center” (Figure 20).

- Put down the large phosphor screen (screen retract, right hand panel). The selected feature should be close by (Figure 21).
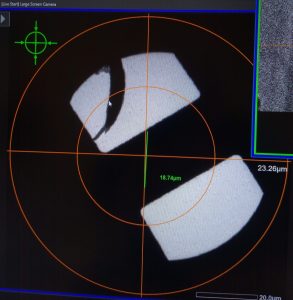
- Use the trackball to move the stage directly on top of the feature you selected on the Atlas image (Figure 22).
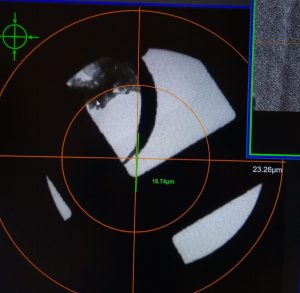
- Re-Insert CL aperture by double clicking on “CL” in the microscope pictogram in the “TEM center” (Figure 23).
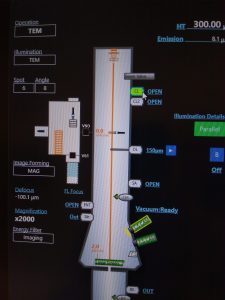
- In Serial EM click on View (dark green panel) to make a single view image with the camera (Figure 24).
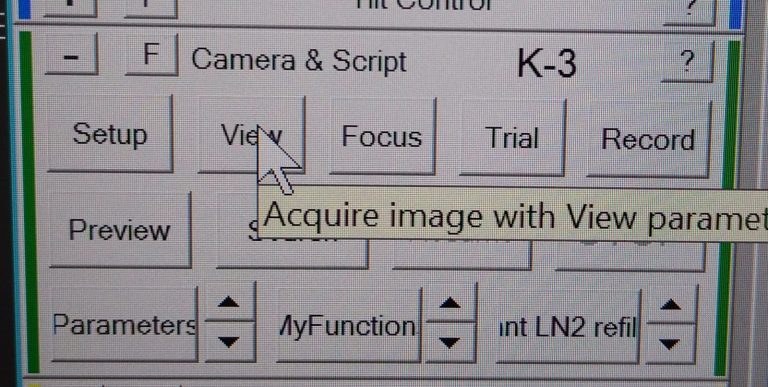
- On the View image mark the same part of your feature you selected on the atlas with a left click. A green cross will appear (Figure 25).
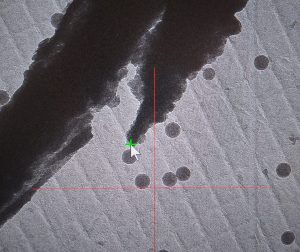
- Ensure that the marker point is still highlighted in the navigator window on click in the Menu bar on “Navigator –> Shift to Marker” (Figure 26).
- In the new pop up window you can see the shift that will be applied on the atlas map. If the numbers are completely different than the amount you used in the MoveStage command you can update the Move stage X Y with more accurate values. Confirm the shift by clicking “OK” (Figure 27). Note: Make sure that the displayed shift amount is reasonable otherwise you might have done something wrong. i.e. not align to the correct feature. Note 2: This new system would allow you to selectively shift only grid atlas point, square montage points or points from a different registration (If needed).
- Save the navigator file by clicking in the menu on “Navigator –> Save” (Figure 28).
- Save the log file by clicking in the menu on “File –> Save Log As…” (Figure 29).
- Ensure that yourcorrect working directory is selected (still under X:) and define the “File name” as “log”. Click “Save” (Figure 30).

- Insert the next grid by highlighting the sample in the “CryoSpecPorter” window and clicking on “Load” (Figure 31).
- Repeat the procedure from Step 2 for all other grids you inserted. Once all grids are done continue with next step.
Perform test shots
- Insert the grid in which you want to do test shots by highlighting the corresponding grid in the CryoSpecporter and click on load. (Figure 1) Note: If there is another grid still on the stage it will be automatically unloaded.

- Open the corresponding navigator file via the Menu bar Navigator --> Read and open (Figure 2).
- Select the correct navigator.nav file in your grids folder and click "open" (Figure 3).
- Open the old log file for appending information via the Menu bar File --> Read & Append (Figure 4)

- Select the correct and corresponding log file (Figure 5).
- In the navigator window double click on the item corresponding to your atlas. It is a "Blu" color item (Figure 6).
- In the navigator window click on "Add Points" and make a left click on all squares that you want to try test shots on. The points appear as red crosses in the image and "red" color item in the navigator window (Figure 7).
- Once finished click in the navigator window on "stop adding" (Figure 8).
- In the navigator window highlight one of your points and click on Go to XYZ (Figure 9).
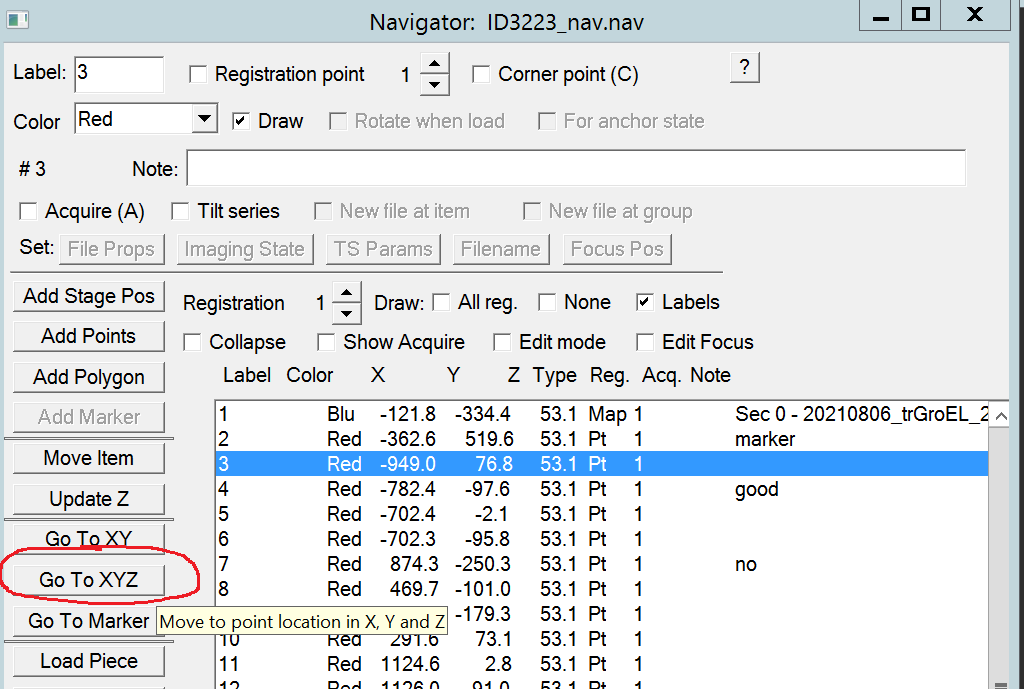
- Measure and go to eucentric height by clicking in the Menu bar on “Task –> Eucentricity --> Rough Eucentricity” (Figure 10).

- When the procedure is finished the log file will say "Rough eucentricity ..... finished.
- Update the z-height of your square point by highlighting the point in the navigator window and clicking "Update Z" (Figure 11).
- Open Camera Setup (dark green panel, Setup) (Figure 12).
- In the View panel select "continuous" for "Acquisition" (Figure 13).
- In the record panel go to "Set folder" (Figure 14). Ensure that "Save frames" is checked. Note: Feel free here to change exposure time and Frame time, but you mostly don't want to.
- Ensure that you are in the correct record folder on (X:) (Figure 15). Click OK.
- In the record panel go to "SetFile options" (Figure 14).
- Change your root file name and click "Ok" (Figure 16).
- Back in the setup window conform all your changes with "OK" (Figure 14)
- Change the parameters of the single hole record by clicking in the Menu bar on Script --> Edit --> Edit 4 (or similar) (Figure 17).
- In the "SingleHoleRecord" script you can adjust the defocus value (DEFOC). the number of shots around the center (NShots), the radius around the center (RAD) and if you want to measure ice thickness (Figure 18). Conform with "OK". Note: Usual values for 2um holes is 5 shots, 0.67 rad; 1.2 um holes is 3 shots; 0.45 rad; 0.6 um holes 1 shot, 0 rad.
- For safety reason in case of crashes save the setting via the Menu bar Settings --> Save (Figure 19).

- Make a continuous view image by clicking "View" in the dark green camera panel (Figure 20).
- Move with the trackball over the hole of interest (Figure 21). The center is marked by a giant red cross.
- Stop the camera w. "stop" dark green panel (Figure 22).
- If you do not see the FFT panel it is recommended to open it now via the Menu bar Process --> FFT (Figure 23)
- Define the location of Focus mode by clicking on Focus in "Define position or area" pink panel (Figure 24). Note: This and the following steps only need to be changed when you change you sample to a different grid.
- Left mouse click over the neighboring carbon area, where you want to do the focusing (Figure 24). The focus position is indicated in yellow. Note: In rare cases the circle is not visible and you have to do once a view image in "Single image mode" (set under Setup, dark green panel).
- Define the location of Trial mode by clicking on Trial in "Define position or area" pink panel (Figure 25).
- Left mouse click over the neighboring carbon area, where you want to do the trial image (Indicated with a red circle) (Figure 25). Note: Focus and trial will always have the same relative orientation.
- Select "None" in "Define position or area" pink panel (Figure 26).
- In the script panel click on "SingleHoleRecord" to do the test shots (Figure 27). Important: If the hole is centered in view mode but the record images hit the carbon area, you need to do a view vs record alignment!
- Repeat Go to XYZ, eucentric rough (Steps 9-12) and singleHoleRecord (last step) on multiple places on the grid until you know if you have a good grid for data collection or not.
- When finished with test shots, save the navigator file (Navigator --> Save) and save the log file (File --> Save log).
- Repeat the whole procedure for all other grids you want to test shoot.
- IMPORTANT: Once you are done with your test shots, set view mode back to single image acquisition (Setup, dark green panel --> View panel --> Acquisition, Single Image)
Square montaging
At this point you have figured out which grid you want to use for automated data collection over night.
Initial preparation
In case you do not use the same grid for data collection than you used just now for test shots until now
- IMPORTANT: Make sure that view mode is set back to single image acquisition (Setup, dark green panel --> View panel --> Acquisition, Single Image)
- Insert the grid in which you want to do test shots by highlighting the corresponding grid in the CryoSpecporter and click on load. (Figure 1) Note: If there is another grid still on the stage it will be automatically unloaded.

- Open the corresponding navigator file via the Menu bar Navigator --> Read and open (Figure 2).

- Select the correct navigator.nav file in your grids folder and click "open" (Figure 3).
- Open the old log file for appending information via the Menu bar File --> Read & Append (Figure 4)

- Select the correct and corresponding log file (Figure 5).
Defining a polygon (Square area)
- Ensure that low dose mode is switched on (pink panel) (Figure 6).
- If not yet done, display the atlas image by double clicking on atlas file in navigator (Figure 7).
- In navigator window select “Add polygon” (Figure 8).
- Mark the 4 edges of a square by clicking on the atlas image. Green points/lines will appear (Figure 9).
- In navigator window click “Stop adding” (Figure 9).
Setting up polygon montage
- Ensure that in the navigator window the previously defined polygon is highlighted. (Green point named Poly1) (Figure 10).
- In the Menu bar select Navigator –> Montaging & Grids –> Setup Polygon Montage (Figure 11).
- Confirm the two next conformation windows with "Yes" and "Ok" respectively (Figure 12&13).
- The montage setup will show you Magnification (2000) and Binning (2), as well as the expected number of pieces. Confirm with “OK” (Figure 13). Note: Only use continuous mode for faster collection (with a settling factor of 0.5) if you are really ok with the additional dose on your sample.
- Confirm the file properties by clicking “OK” (Figure 14).
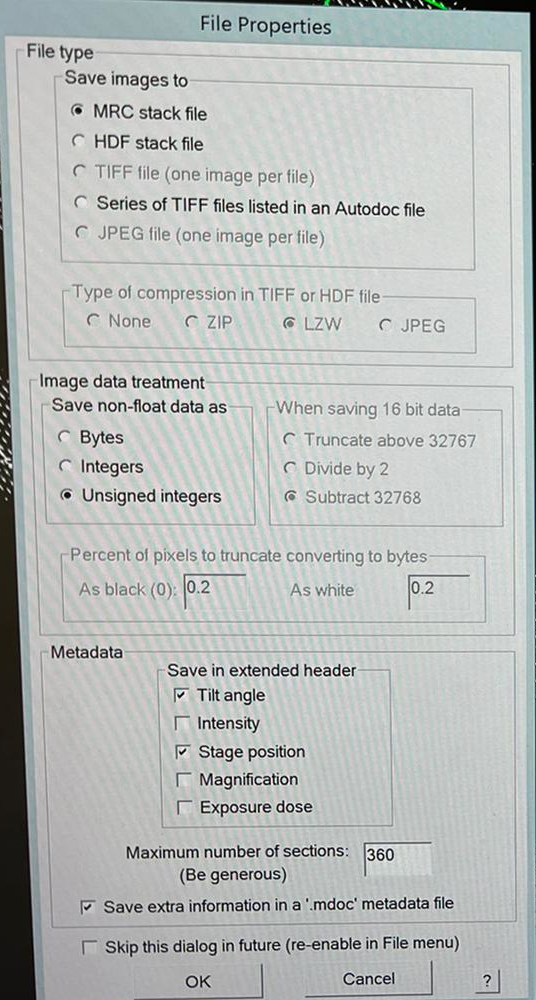
- Ensure to save in your working directory (X:\SerialEM_rawdata\MyGroup\MyName\MyProject\). Call the file name “squares.mrc”. Click “Save” (Figure 15).
Create square images
- Ensure that the Atlas image is shown on the screen. (Double click on atlas file in navigator window to open it) (Figure 16).
- In the navigator window click “Add points” (Figure 17).
- Select in the atlas picture all the squares you want to image. Ensure that you pick the square more or less in the center (Figure 18).
- Click “stop Adding” in navigator window (Figure 18).
- In navigator window mark all square points (Red point) for acquisition by pressing the button “A” on every point. Note: Alternatively you can (un)check “Collapse groups” and select all targets added in one go (Figure 19).
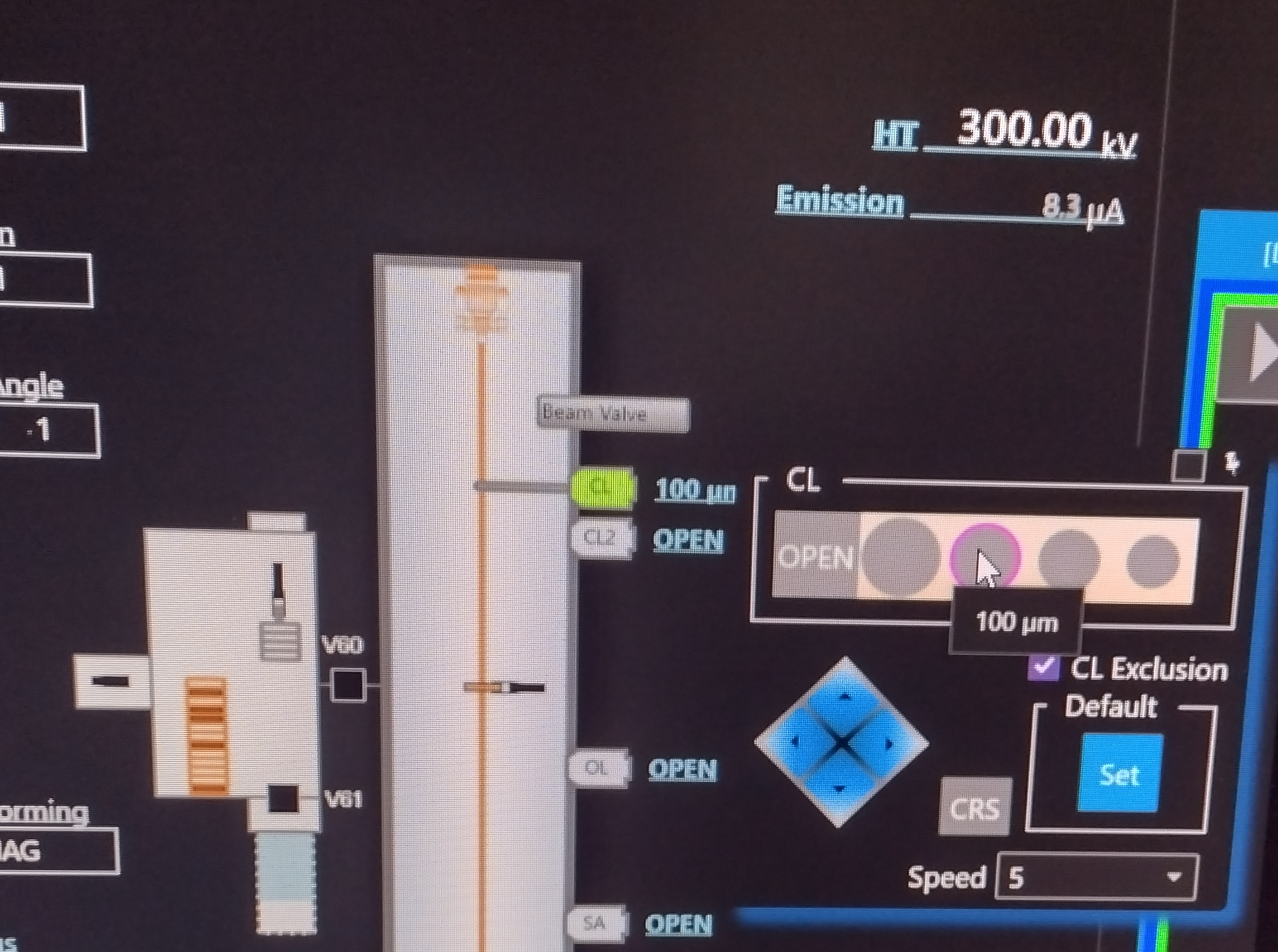
- Change the CL aperture to 150 um by hovering in the TEM center over the "100 um" next to "CL" and selecting the left most aperture which is 150 um (Figure 20).
- Put down the Large phosphor screen (Right hand panel, Screen retract)
- In SerialEM select view mode (pink panel, Vie) (Figure 21)

- Center the beam using Shift X&Y (both hand panels).
- Lift the phosphor screen (Right hand panel, Screen insert).
- In the Menu bar select Navigator –> Acquire at Items... (Figure 22).
- Be on Parameters for "Mapping" and ensure that “Rough eucentricity every item” is active and “Acquire and save image or montage” as well as "Make Navigator map". Click “Go” (Figure 23). Note: "Hide unselected options" and "Hide unused task" can simplify the view and only show active things.
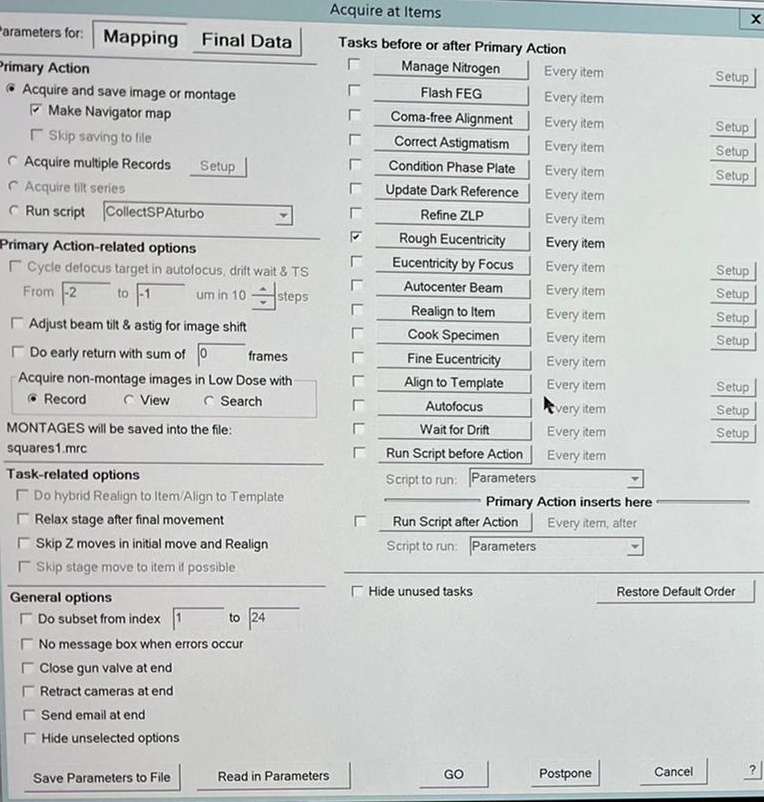
- The microscope will automatically collect a map for each selected square (Red point). The image window shows the current Montaging piece. On the bottom right is indicated which piece is shown (here: 21 of 77) (Figure 24). In the navigator window you can also see the estimated time of completion.
- At the end a confirmation window will appear that shows how many images are acquired (Figure 25). Each square map can be opened in the navigator window with a double left click (as blue point).
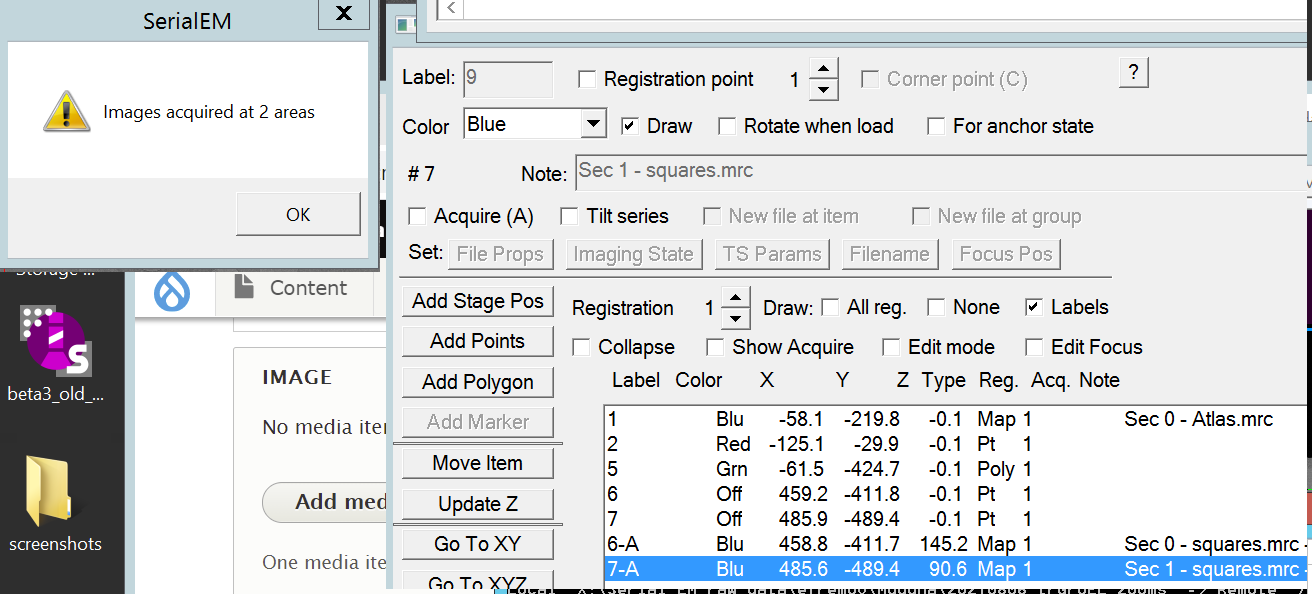
- Re-select the 100 um CL aperture, by hovering in the TEM center over the "150 um" next to "CL" and selecting the second from left aperture which is 100 um (Figure 26).

Define multi hole pattern
This procedure is only necessary if you collect with a multi hole patter. I.e. collecting multiple holes with one stage motion.
- In the navigator window click on "Add Points" and make a left click on all square that can be used for alignments only (since we will burn large areas of the square). The points appears as red crosses in the image and "red" color item in the navigator window (Figure 1). Note: You can add the note in the navigator for this point that is is for alignment.

- In the navigator window on "stop adding" (Figure 2).
- In the navigator window highlight the point corresponding to your alignment square and click on Go to XYZ (Figure 3).

- Measure and go to eucentric height by clicking in the Menu bar on Task --> Eucentric rough (Figure 4).
- When the procedure is finished the log file will say "Rough eucentricity ..... finished.
- Update the z-height of your square point by highlighting the point in the navigator window and clicking "Update Z" (Figure 5).
-
Put down the large phosphor screen (Screen retract, right hand panel)
-
Ensure that you are in view mode conditions (pink panel, Vie.) (Figure 6)
- Remove the CL aperture with a double click on CL (TEM center, microscope pictogram) (Figure 7)
- Use the track ball to move towards the center of the square (We need some space for this procedure)
- Go to Record mode in SerialEM (pink panel, Rec.)(Figure 8).
- Center yourself perfectly over the center of a hole using the track ball (Figure 9). Note: You can change FL focus (counter-clockwise, right hand panel) to increase the viewing area. It does not harm to see a small back circle in the center.
- In the Menu bar go to Navigator --> Montaging & Grid --> Set Multi-shot Parameters (Figure 10).
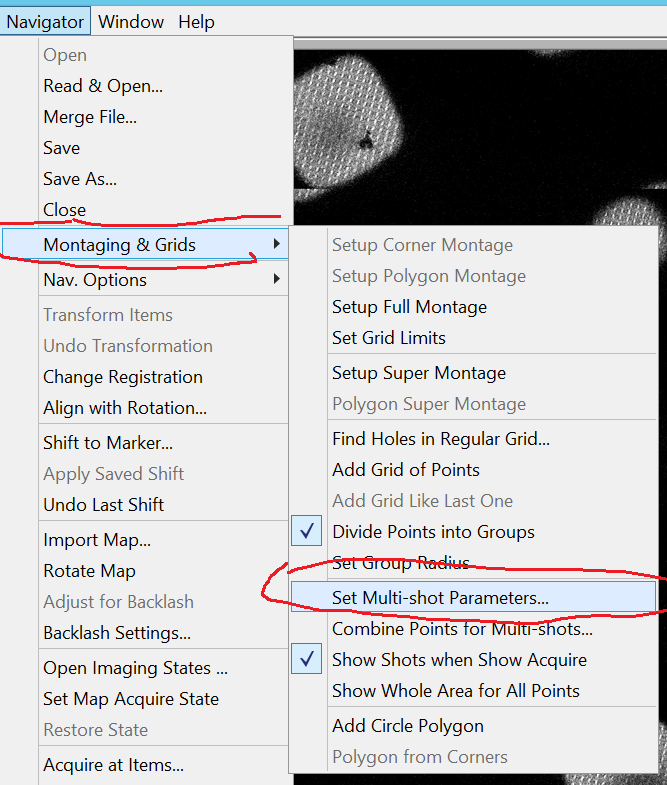
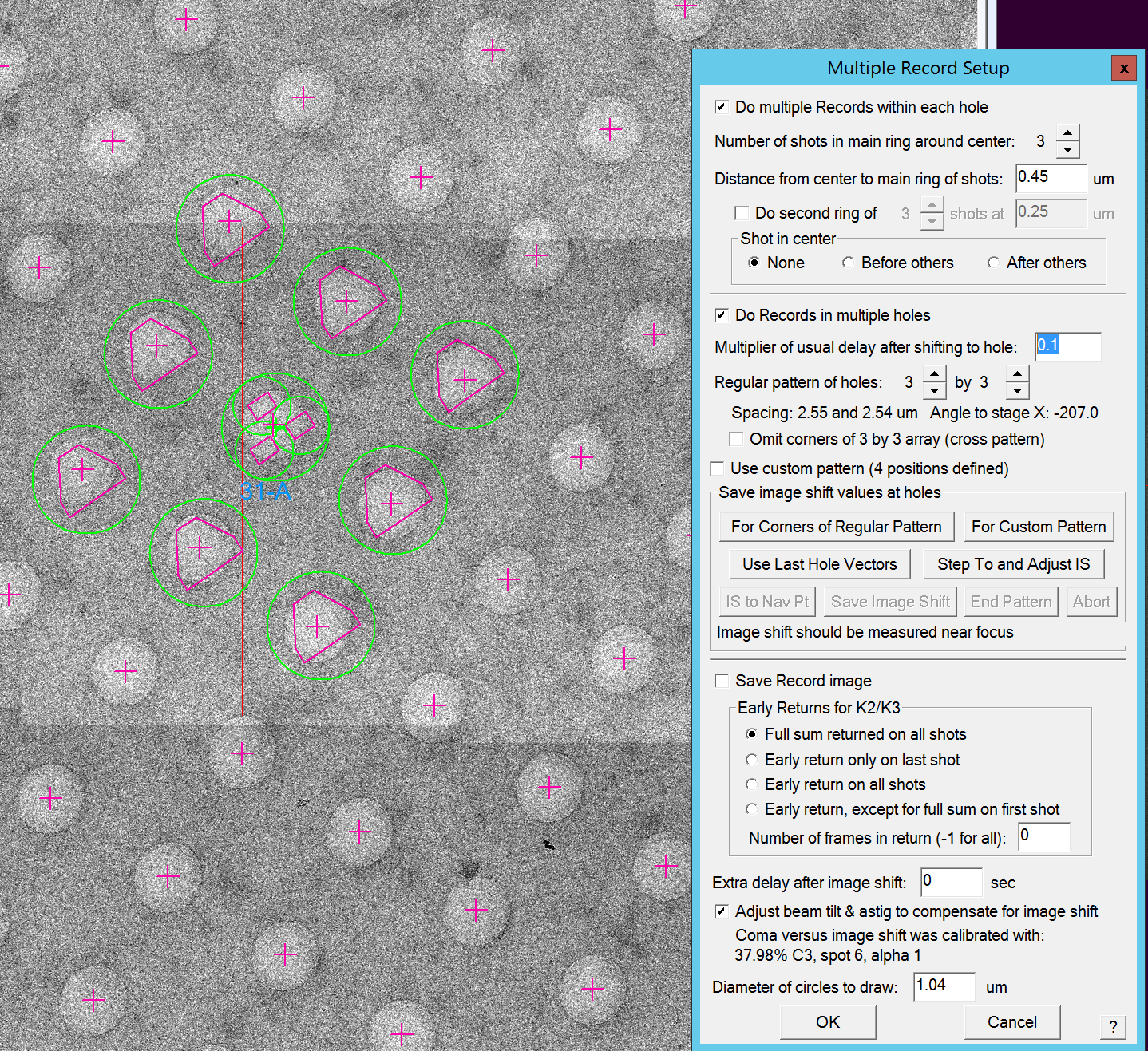
- In the Multiple Record Setup window (Figure 11), make sure that the checkbox "Do Records in multiple holes" is active
- Select the regular pattern of holes that you want to collect (i.e. 3 by 3)
- Start the hole pattern procedure by clicking the "For corners of Regular Pattern" button.
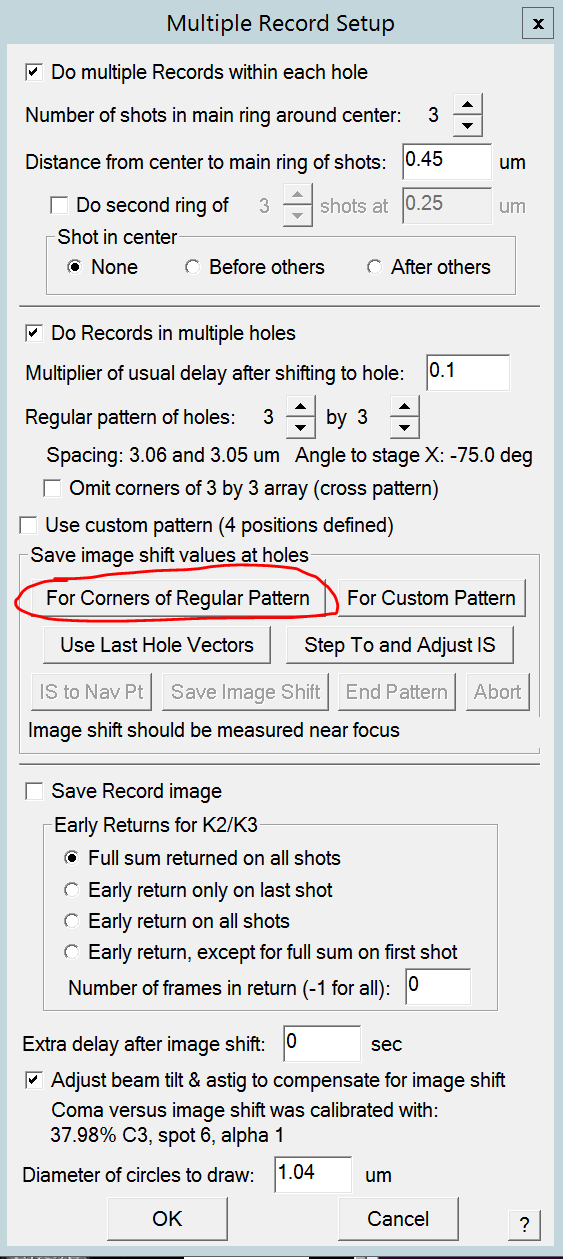
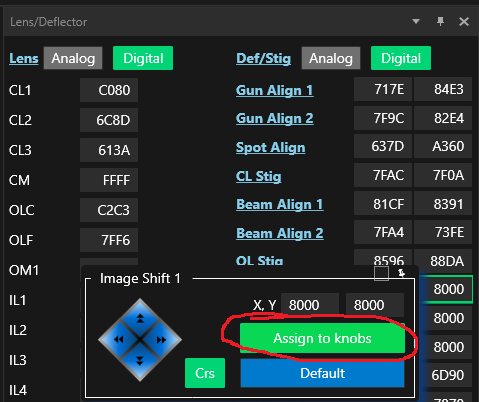
- Assign Image shift 1 to the deflector knobs by hovering in the "Lens/Deflector" window (TEM center) over "Image Shift 1" and click on Assign to knobs (Figure 12). Field will become green.
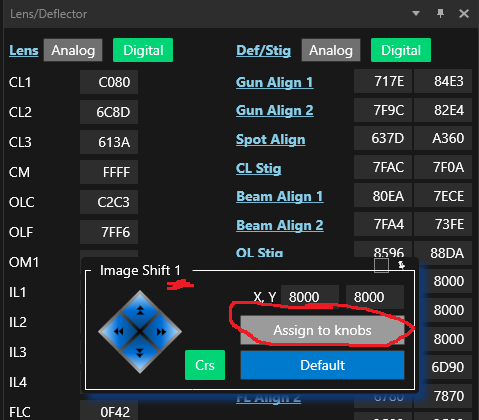
- If needed, you can use Def/Stig X&Y (both hand panels) to more accurately center on the current hole on the phosphor screen. Note: Do not move with the trackball, this will crash the procedure.
- Click "Save Image Shift" button (Figure 13)
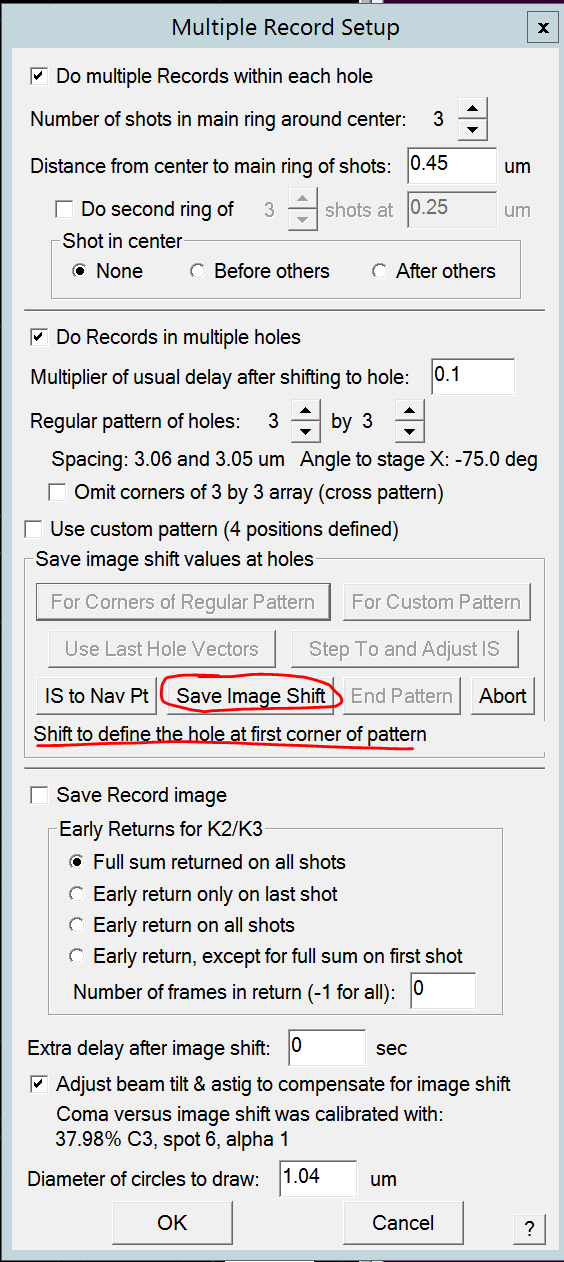
- Use Def/Stig X&Y (both hand panels) and shift in the first direction (any of the four nearest hole directions) SerialEM tells you how many holes you need to shift (i.e. 2 holes) (Figure 14). Note: The amount of holes that need to be shifted is dependent on the hole pattern that you selected. Note2: You will need to adjust the beam center throughout the procedure using shift X&Y (both hand panels).
- Once you are perfectly centered over that hole click the "Save Image Shift" button (Figure 14).

- Use Def/Stig X&Y (both hand panels) and shift in the second direction (perpendicular to the first direction). SerialEM tells you how many holes you need to shift (i.e. 2 holes) (Figure 15).
- Once you are perfectly centered over that hole click the "Save Image Shift" button (Figure 15).
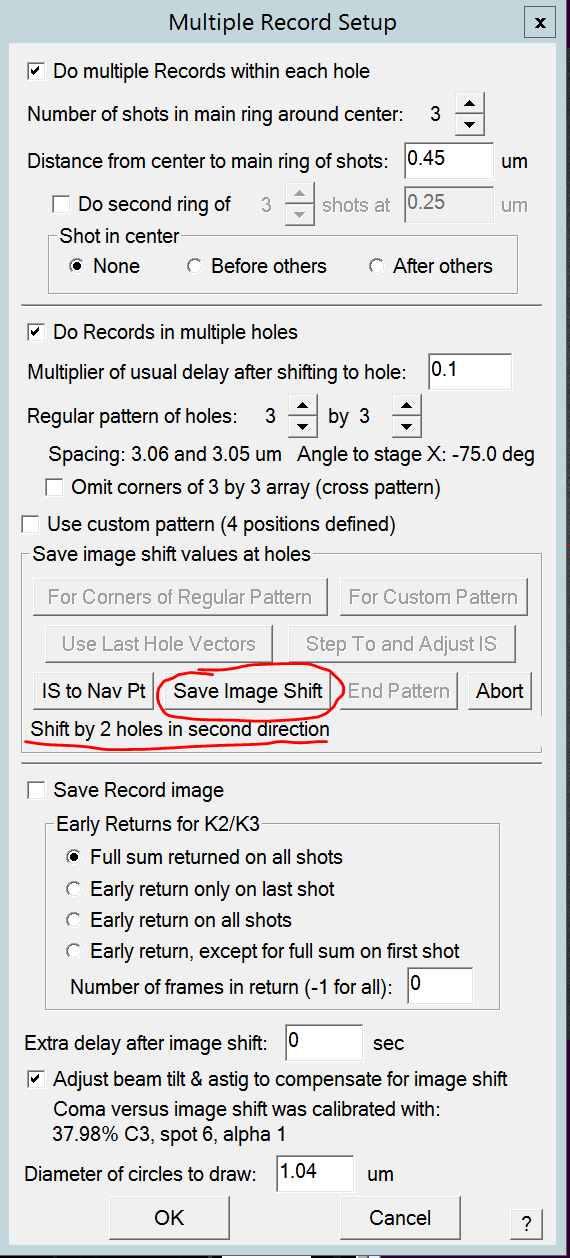
- Use Def/Stig X&Y (both hand panels) and shift backwards in relation to the first direction (180 degrees compared to the first direction). SerialEM tells you how many holes you need to shift (i.e. 2 holes) (Figure 16).
- Once you are perfectly centered over that hole click the "Save Image Shift" button (Figure 16).
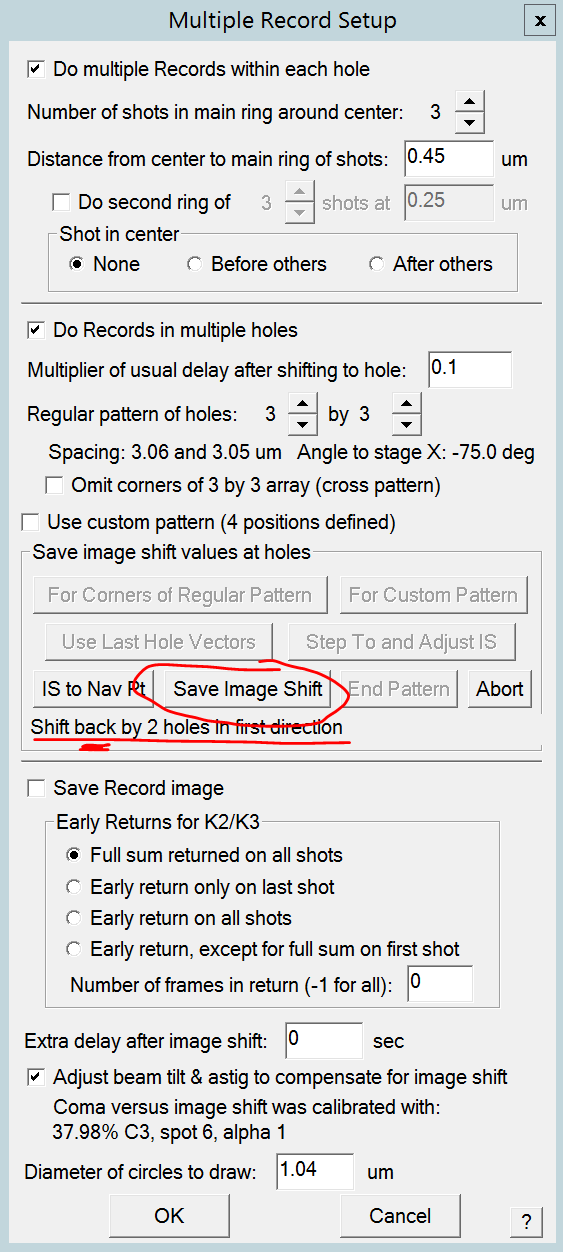
- Optional: You can confirm that the pattern you created makes sense, by left clicking onto a view image and in the center of a hole. The shown pattern should correspond with your actual hole pattern (Figure 17) Note: It will never look perfect because of calibration offsets towards the view mode magnification.
- Click the "OK" button to close the Multiple Record Setup dialogue.
- Re-Insert CL aperture with a double click on CL (TEM center, microscope pictogram) (Figure 18)
- Roughly center the beam, by hovering in the “Lens/Deflector” window over “Beam Align 1” and click on “Default” (Figure 19).

- Center the beam more properly on the large phosphor screen using Shift X&Y (both hand panels)
- Un-assign Image Shift 1 from the hand panels by hovering in the "Lens/Deflector" window (TEM center) over "Image Shift 1" and click on Assign to knobs (Figure 20). Field will become gray again.
- Re-adjust FL focus (right hand panel). Turn FL focus clockwise until the shadow appears from the outside. Turn FL focus counter-clockwise until the beam is again perfectly circular. Turn FL focus dial another 6 clicks counter-clockwise.
- For safety reason in case of crashes save the settings via the Menu bar Settings --> Save (Figure 21).

Finding and combining holes
- Open the acquired square image by double clicking in the navigator window onto the line of your square ( XXX-A; Blue point; named Map; comment SecX - squares.mrc) (Figure 2). Hint: In some cases it can help to have "Align pieces in overview" checked (dark brown panel) to get a nicer stitching before open the map.
- In the Menu bar select Navigator --> Montaging & Grids --> Find holes in regular grid... (Figure 2).
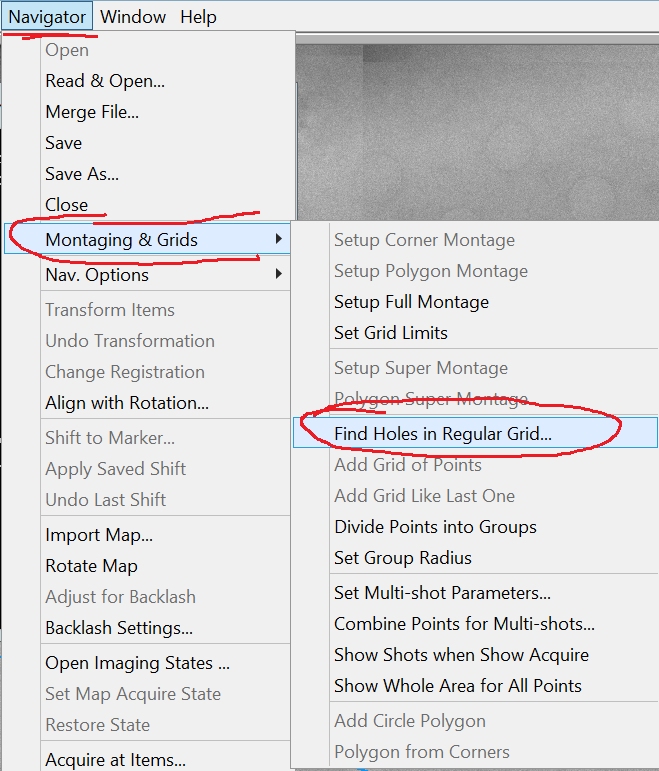
- In the new Hole Finder window specify the hole size of your grid and the periodicity (Figure 3). Note: periodicity is hole size plus distance between holes! Note 2: If your grid hole sizes differ from manufacturer specification you can measure the distances in the view image or square montage by holding the shift button on the keyboard and clicking & dragging the left mouse button on the image.
- The maximum error may need some adjustment and should be changed in a manner that it maximizes picking coverage and accuracy. (We tested successfully up to 0.4 um error).
- Click the "Find holes" button (Figure 3).
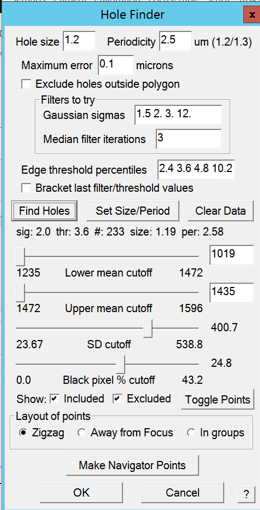
- SerialEM may display a choice dialogue (Figure 4). Click "Go On" to proceed or "Skip analysis" to proceed and not be asked again. Note: Out of experience the autocorrelation peak is always worse than your entered parameters.
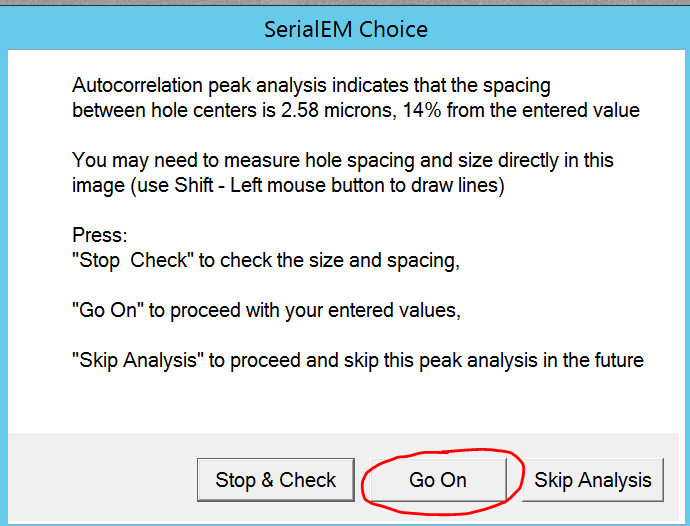
- SerialEM will display all picked holes with a cross. Change the input parameters and click find hole again, if you do not like the picking. Note: The goal is to maximizes picking coverage and accuracy.
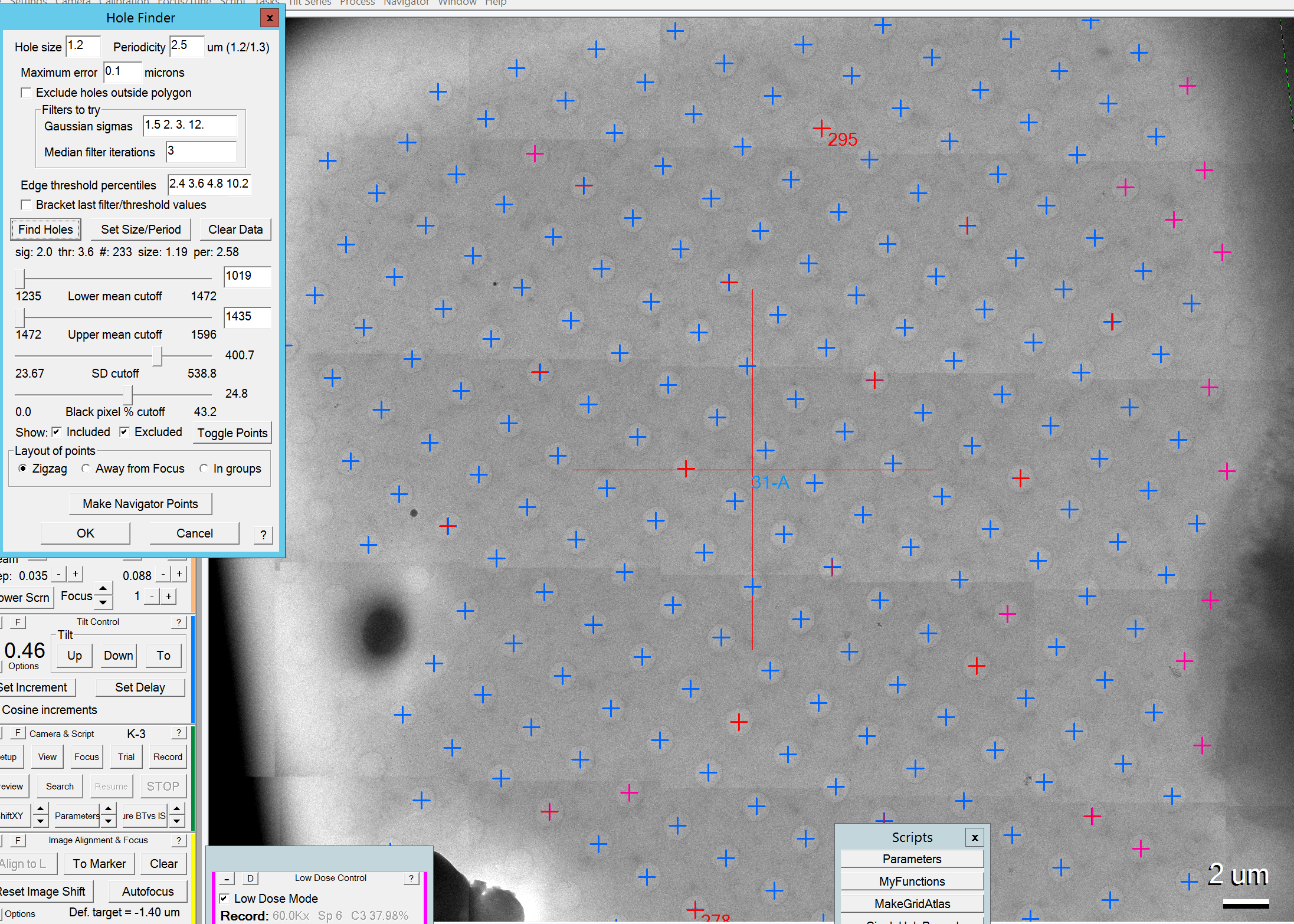
- You can change the four sliders to adjust which holes should be included (pink) or excluded (blue) for data acquisition (Figure 6).
- Once you are happy with the results, click the "Make Navigator Points" button (Figure 6).
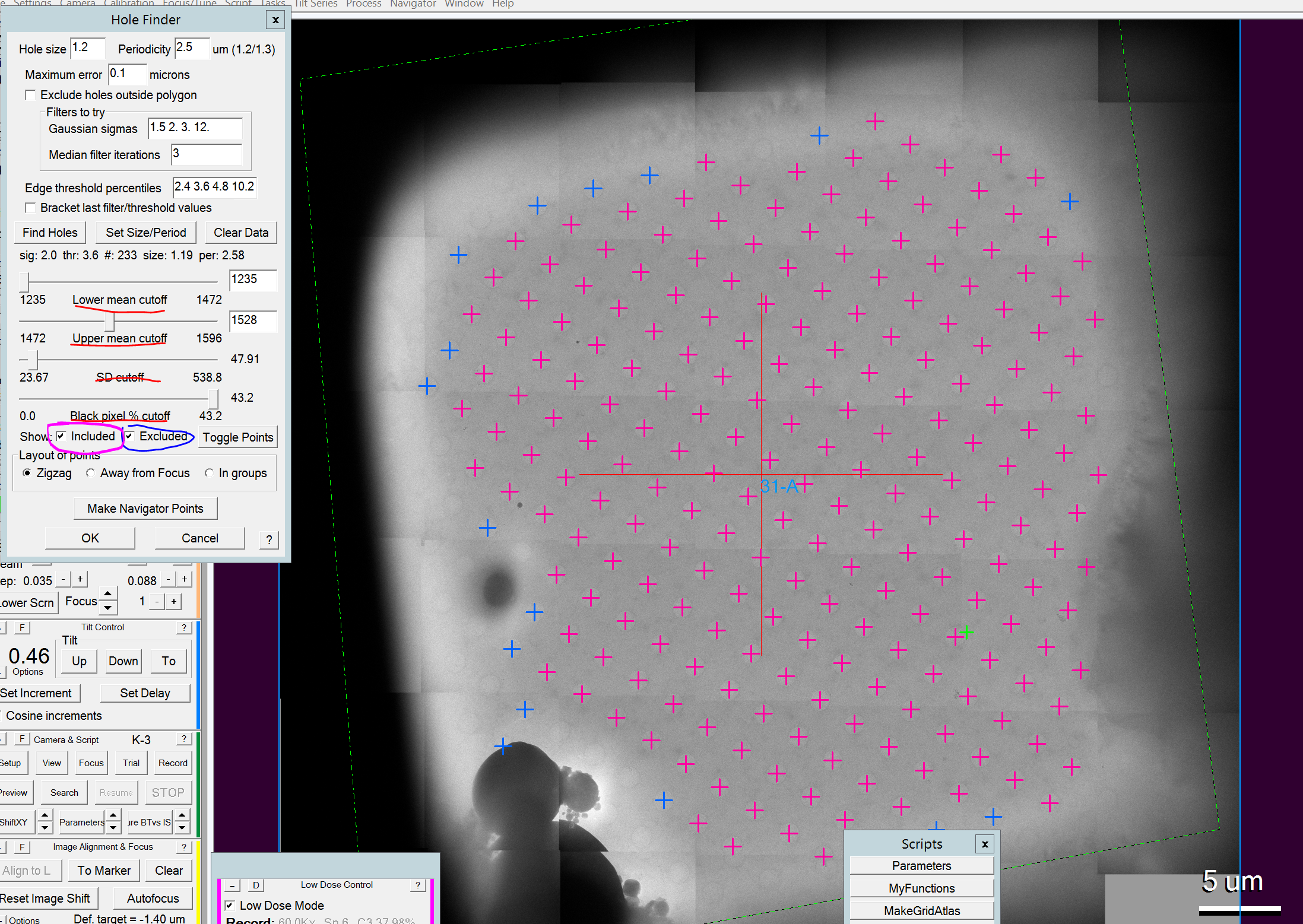
- Now you can still manually delete targets. I.e. those to close to the edge of the square or bad targets that could not be excluded with the slider. You can do this by clicking on one of the hole items in the navigator window (Figure 7) and clicking the delete button in that window. Hint: Use the up and down cursor on the keyboard to quickly flick through the targets.
- Once you are done the square picking might look like Figure 8.

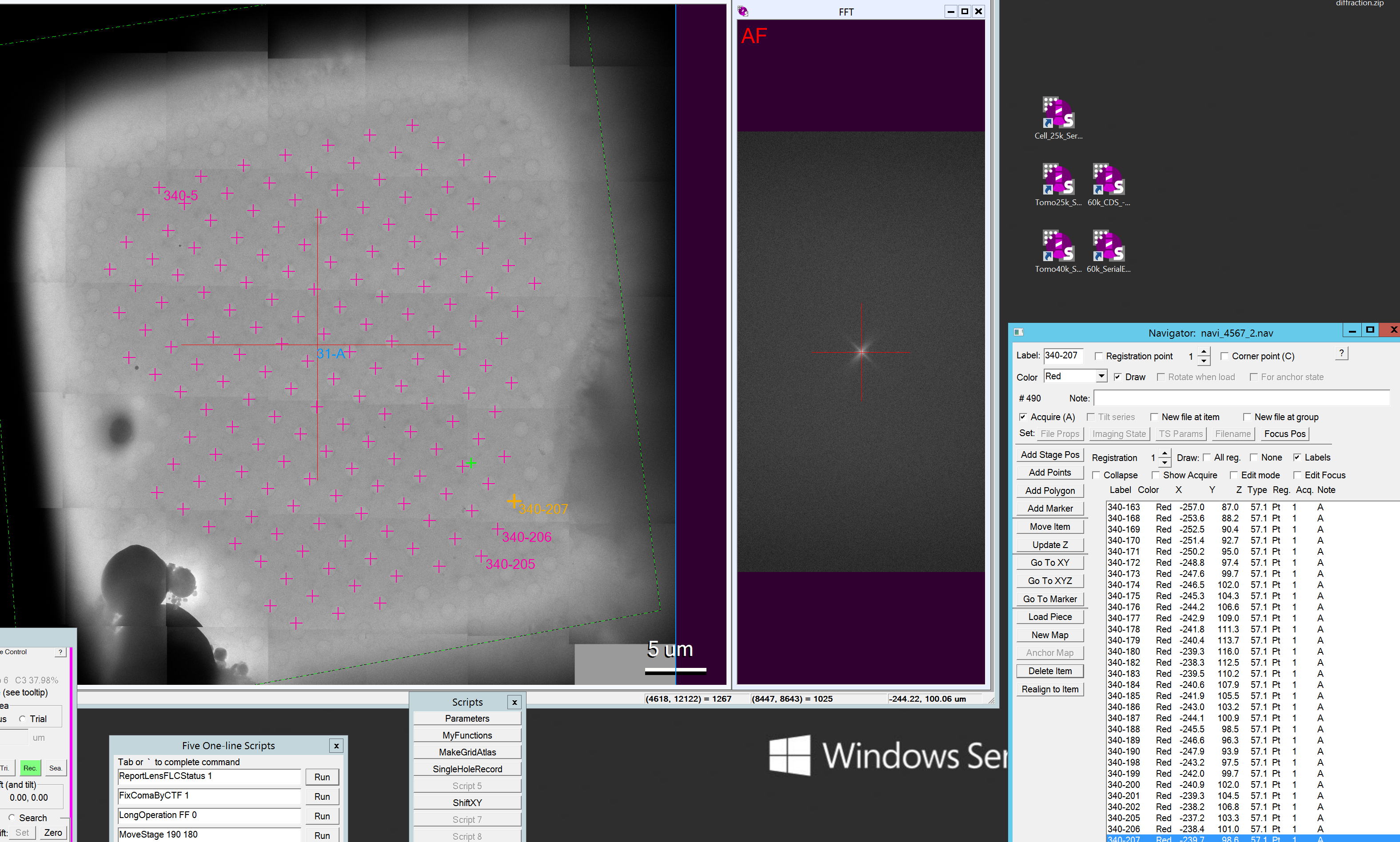
- You now want to combine the holes into acquisition targets for multi hole collection. Important: You have to have done the hole pattern by now and this cannot be done in the dummy. You can pick in the dummy but then need to later combine in the active SerialEM executable!
- In the Menu bar click on Navigator --> Montaging & Grids --> Combine Points for Multi-shots (Figure 9).
- Make sure that you have selected any point in the group of items you want to combine for multi-shot and click in the Multiple Hole Combiner window the "Combine Points" button (Figure 10).
- The list of single items will be gone from the navigator window and is replaced with the Multi-hole shot items (Figure 11)
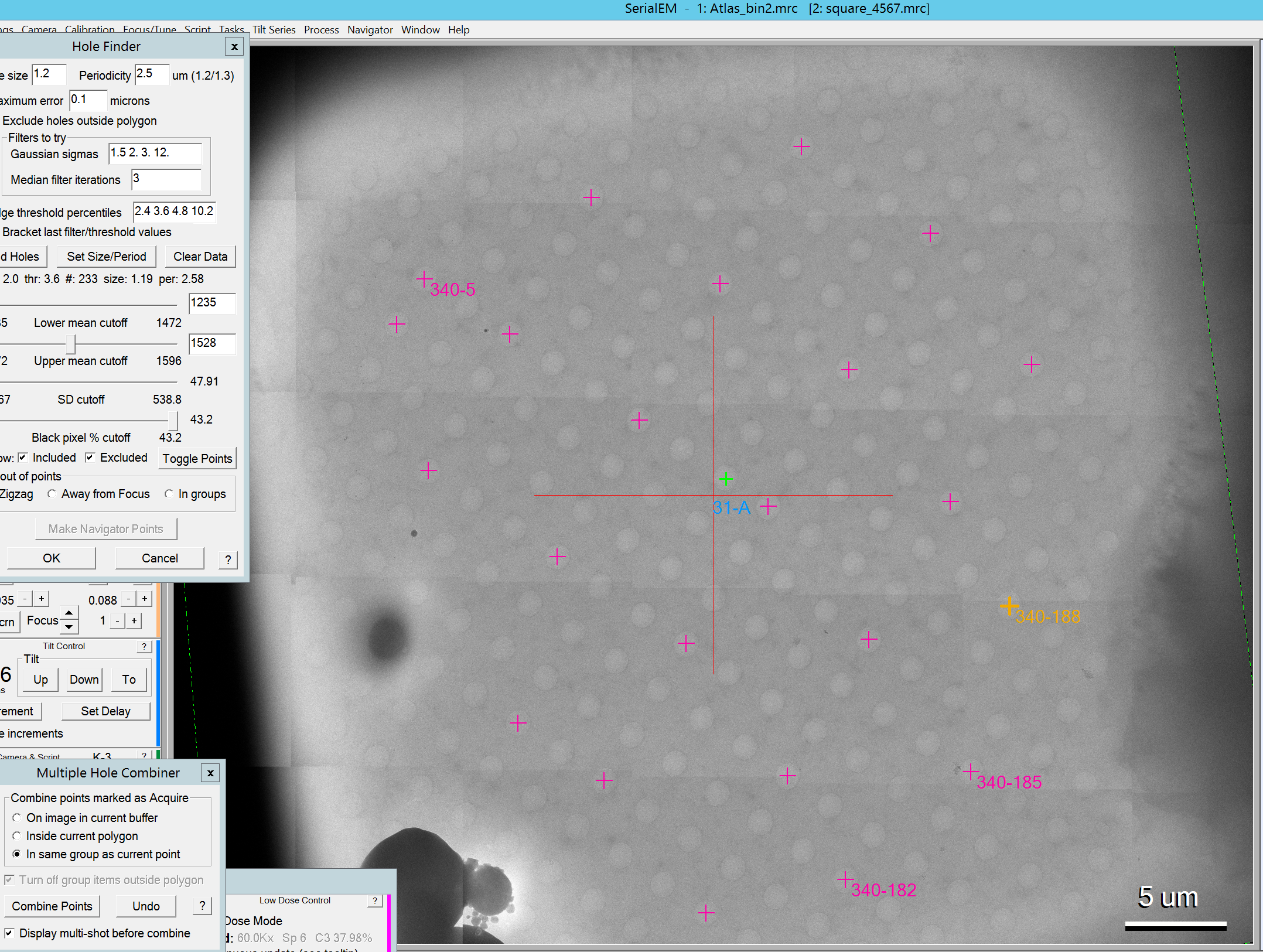
- Optional: You can display which and how many holes are shot for each item if you activate the "show acquire" checkbox in the navigator window and select any of the acquisition targets therein (Figure 12).
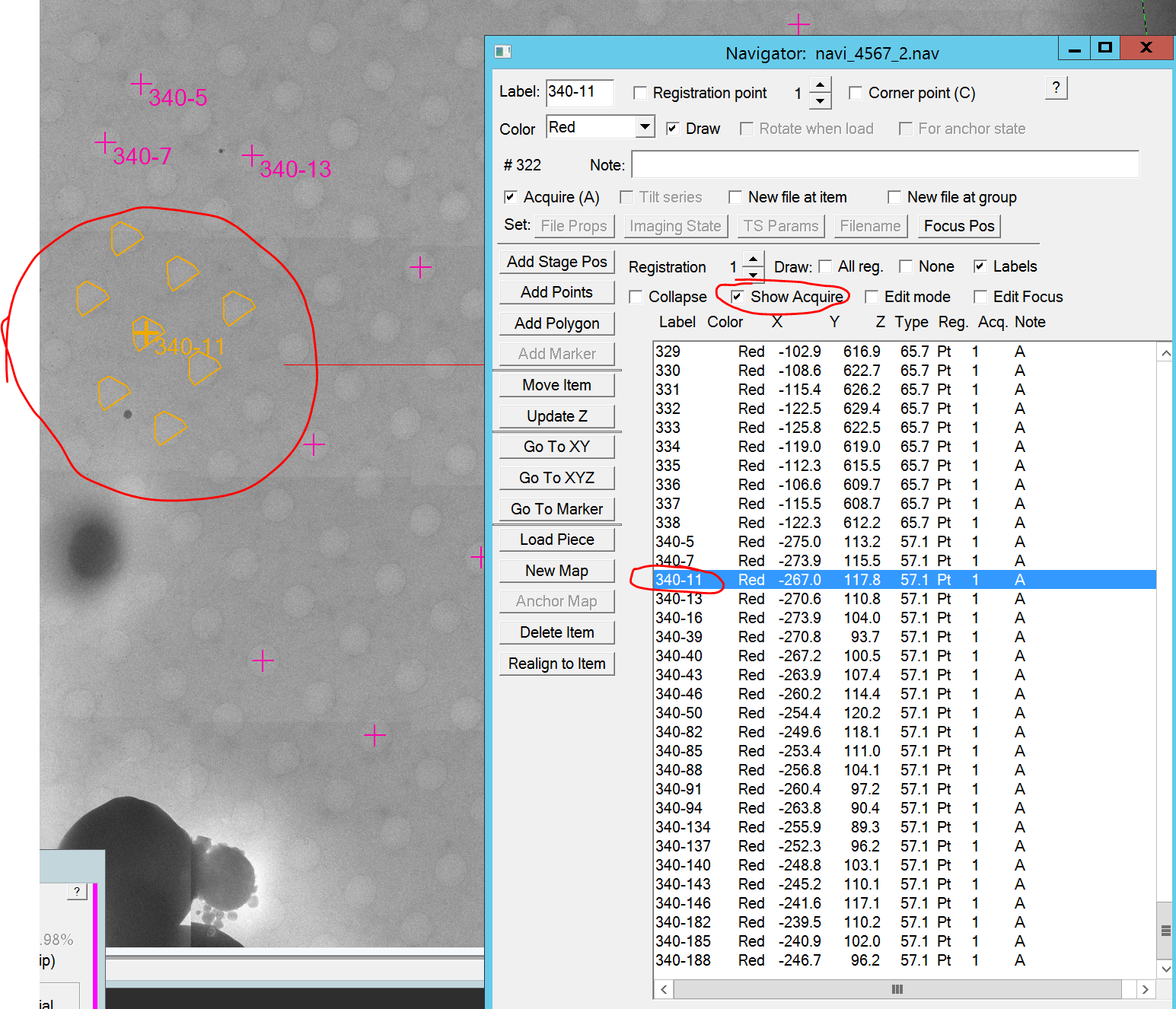
- It is recommended to always save the Navigator between each hole finding, as SerialEM may crash if the picking diverges. Do this by clicking in the Menu bar on Navigator --> Save (Figure 13).

- Repeat hole finding and combining for all your squares. Note: You can first pick and combine only two squares, then start the collection. While it is running you can do the hole finding of the other squares in the dummy, save the navigator file and then combine the hole in the active SerialEM executable (after ending the navigator of course).
Coma free alignment
Make sure you do not have residual drift during the procedure as this will harm the results.
- If not there yet, go to your alignment square by highlight the point corresponding to your alignment square and click on Go to XYZ (Figure 1).
- Important: Make sure that you are actually at eucentric height. (I.e. Do rough eucentricity, go to record mode, hit std focus, measure defocus and manually adjust z-height with the small hand panel on the left). Often This should have already the proper z-height due to procedures before.

- Put down the large phosphor screen (Screen retract, right hand panel).
- Hit Standard Focus (STD Focus, right hand panel).
- Move the stage with the track ball over the carbon area.
- Center beam with Shift X&Y both hand panels.
- Define a target defocus by clicking in the SerialEM Menu bar on Focus/Tune --> Set Target (Figure 2).

- Set the target to -1.3 um defocus (Figure 3).
- In the Menu bar go to Focus/Tune --> Autofocus to do auto-focusing (Figure 4). Hint: If the focus position ends up in the hole and focusing fails, you may want to temporarily set "Define position of area -- Focus" to 0 um offset. Undo this before starting the data collection.

- Do coma free alignment by clicking in the Menu bar on "Focus/Tune --> Coma-free alignment by CTF" (Figure 5). Important: Check in the FFT image that the fit (yellow dashed lines) corresponds to measured location of Thon rings.

- The log file reports the amount of beam tilt correction (Figure 6). Hint: High score and low fit to values indicate a good run.
- Do astigmatism correction by clicking in the Menu bar on "Focus/Tune --> Coma astigmatism by CTF" (Figure 7).

- The log file reports the amount of OL stigmator correction (Figure 8). Hint: High score and low fit to values indicate a good run.
- Repeat steps 9-12 until stable. Sometimes you do 2 or 3 cycles. Important: Long term view mode affects the beam tilt temporarily. Try not to do it directly after hours of square montaging or redo it after 1 h of running data collection.
View vs Record alignment
I assume here that you are still on your alignment square you used to do coma free alignment
- Ensure you are in record mode (Figure 1)
- Ensure that there is no IS currently applied to the microscope (orange panel) (Figure 2)
- If there is an Image shift reset image shift (yellow panel) (Figure 3). If not, continue to next step.
- Put down the large phosphor screen (screen retract, right hand panel).
- Optional: Remove the CL aperture with a double click on CL (TEM center, microscope pictogram) to increase the field of view (Figure 4).
- Find a feature on your (alignment) square that is identifiable in View and record magnification. Center it on the phosphor screen using the track ball (Figure 5).
- Re-Insert the CL aperture with a double click on CL (TEM center, microscope pictogram) (Figure 4).
- Make a single view image with SerialEM (dark green panel, View) (Figure 6).
- If there is a mis-alignment between View and Record mode (i.e. because you adjusted the beam tilt) you will see the feature off centered (Figure 7).
- Keep the right mouse button pushed on the image in SerialEM and drag the feature to the center (indicated with a giant red cross) (Figure 8).
- Confirm the offset in the pink panel by clicking "Set" under "Offset for View" (Figure 9).
- SerialEM will make a new image with the feature now centered (and no grey bars) (Figure 10).
- Go back in Record mode (pink panel, rec) (Figure 11).
- Put down the large phosphor screen (screen retract, right hand panel).
- Confirm that the feature is still centered on the phosphor screen. If not, redo the procedure.
Starting up data collection
Centering CL aperture
- Go to record mode (Figure 1).
- Put down large phosphor screen (screen retract, right hand panel).
- minimize the beam with the brightness dial (left hand panel, counter clockwise) (Figure 2).
- Center beam with Shift X&Y (both hand panel) (Figure 2).
- Spread the beam until more or less the size of the inner orange ring (brightness dial, clockwise, left hand panel) (Figure 3) Note: If you do not see the orange circle, right click on the large screen camera image --> Show circle scale
- If the beam is off-centers now, move the position of the CL aperture until the beam appears centers again by hovering over the CL aperture size value and use the arrows to move it (Figure 4).

- The beam should now be centered again, indicating a centered CL aperture (Figure 5).
Aligning beam shift for different illumination modes
- Go to record mode (Figure 6). Note: This also restores the brightness value which was changed in the last procedure.
- Put down large phosphor screen (screen retract, right hand panel).
- Center beam using shift X&Y (both hand panels).
- Go to view mode (Figure 7).
- Activate the check box "Set" under "additional beam shift (and tilt)" (pink panel) (Figure 8)
- Center beam using shift X&Y (both hand panels).
- Un-check the previous box "Set" under "additional beam shift (and tilt)" (pink panel) (Figure 8). Note: Every time you adjust beam tilt in record and do a view vs record alignment, you only need to change the beam shift offset between record and view and you can skip focus and trial.
- Go to focus mode ("Foc.", pink panel) (Figure 9).
- Note: The beam is expected to be centered. Hit "reset" under "additional beam shift" if the values are not zero (or basically zero) (Figure 4). Important: If you need significant beam shifts here, the CL aperture is not properly centered.
- Activate the check box "Set" under "additional beam shift (and tilt)" (pink panel) (Figure 8)
- Center beam using shift X&Y (both hand panels).
- Un-check the previous box "Set" under "additional beam shift (and tilt)" (pink panel) (Figure 8)
- Go to trial mode ("Tri.", pink panel) (Figure 10).
- Note: The beam is expected to be centered. Hit "reset" under "additional beam shift" if the values are not zero (or basically zero) (Figure 10). Important: If you need significant beam shifts here, the CL aperture is not properly centered.
- Activate the check box "Set" under "additional beam shift (and tilt)" (pink panel) (Figure 9)
- Center beam using shift X&Y (both hand panels). Note: An accurate relative center between trial and record is very important as this ensures a centered beam during data collection.
- Un-check the previous box "Set" under "additional beam shift (and tilt)" (pink panel) (Figure 9)
- Go to record mode (Figure 10).
- Note you may need to cycle again through Rec. --> Foc --> Tri. in case the beam is not stable yet
Confirm camera parameters
- Open Camera Setup (dark green panel, Setup) (Figure 11).
- In the View panel ensure "single image" for "Acquisition" is selected (Figure 12). Note: The values shown are generally used for carbon grids and the K3 in CDS mode
- In the record panel (Figure 13) ensure that "Save frames" is checked. Click on "Set Folder". Note: Feel free here to change exposure time and Frame time, but you mostly don't want to. we have it set to give you approx. 1 e-/A2/frame and 60 e-/A2 total dose
- Ensure that you are in the correct record folder on (X:) and use the folder corresponding to your current grid (Figure 14). Click OK.
- In the record panel go to "SetFile options" (Figure 13).
- Change your root file name to a proper name. This will be the name of your final data. Click "Ok" (Figure 15).
- Back in the setup window confirm all your changes with "OK" (Figure 13).
Confirm general data collection parameters
- Edit the general parameters in script 2 via the menu bar Scripts --> Edit --> Edit2 (Figure 16).
- Change the minimum and maximum defocus limit you want to use during data collection (Figure 17). Note: You want to use as little defocus as possible, while still being able to see and pick particles.
- You can also change the step size (i.e. the increment of defocus change between acquisition targets)
- Confirm all changes by clicking on "OK
Save Settings
- For safety reason in case of crashes save the settings via the Menu bar Settings --> Save (Figure 18).

Start data collection
- Ensure that in the navigator window only hole targets on squares are marked with "A' for acquisition and no points from the grid atlas.
- Have a control look over the z-height of the hole targets. If any are out of the ordinary (larger or smaller than all others) it indicates a faulty eucentric height measurement. Redo eucentric height for those areas and update the z of that group (collapse group --> highlight the group --> update z).
- Select in the menu bar Navigator --> Acquire at Items... (Figure 19).
- Be under "Final Data" tab and make sure that you selected "Acquire multiple records" (left side), "Run Script before action (CollectSPA_internalSerialEM)" (right side) (Figure 20).
- If you want to measure the ice thickness of the acquisition items with the energy filter you also need to select "Run Script After Action (Measure Ice)" (Figure 20). For this you can also decide how often it runs (e.g. every 5th item). Note: To display the details of the "Run Script after action" task, you need to have that option highlighted.
- Ensure that the multiple records parameters are set properly by clicking on the Setup button next to "Acquire multiple records" (Figure 21).
- Make sure that the parameters are correct and you (i) do multiple records within each hole, (ii) have the right number of shots in the hole, (iii) do records in multiple holes and (iv) have the correct regular hole pattern (Figure 21).
- If this is fine click "OK" in the Multiple Record Setup Window (Figure 21).
- Click "Go" in the Acquire at Items dialogue (Figure 21).
- Important: NEVER stop the data collection by hitting STOP in the dark green panel. If you did you need to redo coma free alignment.
- If you want to change any parameters or re-start with an updated navigator file, hit "End Navigator" (dark green panel) (Figure 24) and wait until the current acquisition target loop is finished. The big red STOP button will be gone when finished.
- Note: In the navigator window you can see the number of targets left and the estimated completion time (Figure 25). The values will stabilize after a few rounds.
Unclipping cartridges
Note: Never touch the cartridge with your bare hand. If you touch it use gloves.

- Hold the cartridge with a blunt tweezer on one side (Figure 1). Alternatively: Use your fingers to hold the cartridge and cover one half of the clipring.

- Use an old tweezer (it will have a bend tip, but needs to be thin) and go, via one of the cartridge side grooves, with the tweezer below the clipring (Figure 2). Note: It helps to use the groove which is the furthest away from the open side of the clip ring.

- In a continuous motion slide the tweezer in and grab the clipring, which will jump out fast from the cartridge (Figure 3). Having the finger with gloves half on the clipring can help. Important: These clip rings are expensive and reusable. Do not loose them.
- Use the cartridges for clipping new grids after drying or store them in the small glass vessels.
Bake out
1) Stop the current collection in SerialEM.
2) Save the Navigator and Log file of the current run.
3) Close SerialEM.
4) Put the Grating grid onto the stage using the Cryospectporter window and the command “Load”.
5) Remove all Cartridges from the Sample Storage.
6) Switch off Emission by hovering over “Emission” in the TEM center microscope pictogram and click in the line Auto Emission “OFF”.
7) Wait until emission reached 0 uA.
8) Switch off High Tension by clicking “HT” in the TEM center microscope pictogram and click in the line for HT on “OFF”.
9) Wait until you reached 0 kv high tension.
10) In the TEM center open the window “LN2 Auto Refill Maintenance”.
11) Click on “Heater Controller”.
12) In the new window select the finish date and time (normally Monday morning, 7 am).
13) Ensure that the Cooling time is set to 8 hours.
14) Ensure that the all targets are selected (i.e. Column, Sample Storage and Stage).
15) Click in the section “Bake Out operation” on “Start”.
16) In the pop_up window confirm with “Yes”.
17) The bakeout will last until the selected time.
18) In Digital Micrograph select in the menu bar “Camera –> Temperature” (Power User mode must be enabled).
19) In the confirmation window select “Yes”.
20) In the next window select a target temperature of 50 degrees celsius. Click “OK”.
21) In the next window click on “Options…”.
22) Select a Heating duration of 8 hours and a Start delay of 1 hour.
23) Confirm with “Ok”.
24) Click on “Start” to run the heating cycle of the camera.
Common problems
Atlas problems
Problem: If I move to a new position to control the shift, I end up somewhere else.
Solution: Move back to the original feature with the navigator item. Confirm that the feature is really the one selected in the atlas image. IF not, redo view vs. atlas alignment
Problem: I definitely aligned the right feature in view mode vs the atlas. But all other points do not fit. (Very unlikely)
Solution: Recalibrate stage movement in 50 times magnification (done by staff only).
Change Liquid nitrogen tank
The full tank last around 8 days, as long as the stage and sample storage tank are filled always simultaneaously.
This happens automatically during the data collection. But if the scope is idle you can use the script "Constant LN2 refill check" in SerialEM scripts panel.
The tank has a level indicator to show how much LN2 is left.
- Check if the tanks are being refilled at the moment (Icon next to microscope pictogram, TEM center) (Figure 1). Wait if so.
- If the tanks are indicating less than 50 % it is worthwhile to trigger a manual refill. If not, continue step 5.
- Open the LN2 Auto Refill Maintenance window (right click on microscope pictogram --> LN2 Auto Refill Maintenance) (Figure 2).

- Click in the LN2 Auto Refill Maintenance widow on "Refill" for both tanks and wait until refilling is finished (Figure 3).
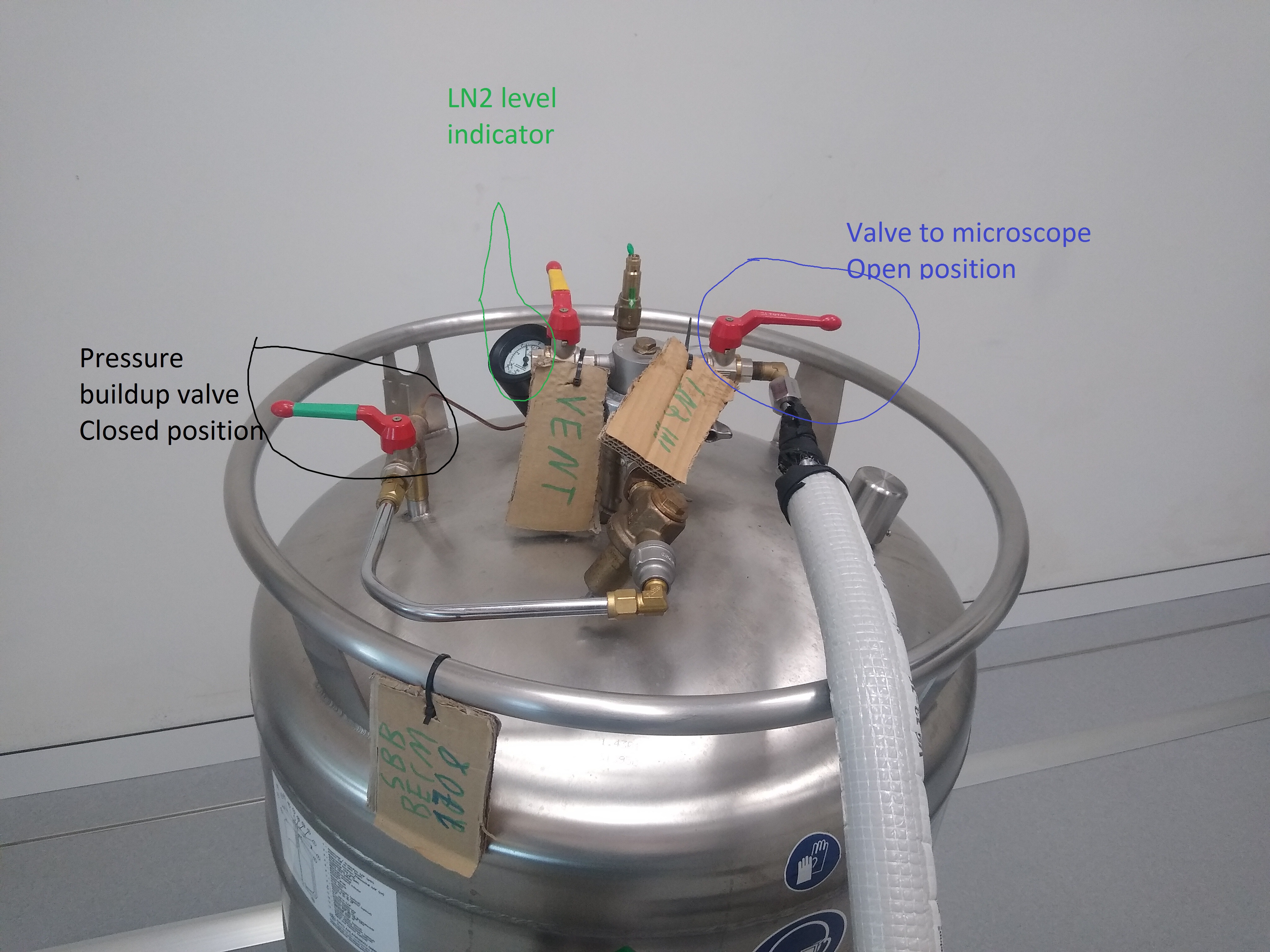
- Enter the microscope room and warm up the hose with the hair dryer, if the connection is icy from a recent refill (Figure 4).
- Close the tank valve leading to the microscope (Figure 4).
- Unscrew the hose leading to the microscope at the tank side (Figure 4).
- Bring the 200 l tank into the technical room to be emptied completely before bringing the tank to the magazine for refill (Figure 4).
- The new full tank should be attached with the hose to the microscope (Figure 4). Make sure the screw is tightened with the wrench. Important: Do not over-tighten!!!
- Open the tank valve leading to the microscope (Figure 4).
- Ensure that the pressure buildup valve is closed (Figure 4).

Gatan Camera crashes and recovery
Sometimes Gatan can crash. This may be either just a software crash (currently most likely) or an actual hardware crash. Both are easy to solve. You can add your email to a script to be informed by email about DM crashes automatically.
Software crash
If DM (Digital Micrograph) crashed, it's icon is missing from the bottom bar and SerialEM will have stalled with an error message
- Do not yet touch SerialEM.
- Re-open DM by clicking on it's software icon on the Desktop (Figure 1).

- Be patient, as it takes a few moments.
- Once the program is up and controllable open the panel of "Camera monitor" on the left hand side (Figure 2).
- Wait until the light of the "Health Status" turns green (Figure 2). Note: If it will never turn green and goes red, you have a hardware crash/problem of the camera.
- Once the light is green you can go back to SerialEM and confirm the error message with "OK". SerialEM will remain in a Paused state.
- If the beam valve closed in the meanwhile, re-open the beam valve (left hand panel).
- In SerialEM just click on "Resume N" (Resume Navigator) and the collection will just continue (Figure 3). Note: It will take some time for the camera to be inserted, don't panic!
Automatic Emailing for DM software crashes
- On the Camera PC go to the "Documents folder" (Figure 1).
- Open the file DMcrash.ps1 with a text editor (right mouse button) (Figure 1).
- In line 8 add your email address (comma separated and in quote marks) (Figure 2).
- Save the file (File-->Save) and close the window.
Hardware Restart Gatan K3
- Before you assume that it is a hardware problem, ensure that it may not be just a software issue.
- Therefore, test a restart of the Gatan DM software as described above first.
- If the software restart did not work, close DM3 and close SerialEM (Save navigator, settings and log first).
- Wait 1 minute for the Camera to retract fully.
- Shut down the camera PC.
- Go into the technical room and shut down the Camera Power supply, by touching the large G-icon (Power supply is located on the top of the left rack).
- Wait 1 minute.
- Restart the Camera power supply (Power supply is located on the top of the left rack).
- Wait 5 minutes.
- Restart the Camera Computer (The computer is located on the top of the rack on the back left)
- Log in to the System (password is written on the physical PC in the technical room)
- Open DM by clicking on the Deskop icon (Figure 1).

- Be patient, as it takes a few moments.
- Once the program is up and controllable open the panel of "Camera monitor" on the left hand side (Figure 2).
- Wait until the camera cooled stably back to -20 degrees and the light of the "Health Status" turns green (Figure 2). Note: If it will never turn green and goes red, you have to repeat above procedure.
- Make one test image to ensure the camera is reacting properly (either using SerialEM or DM itself). Note: It will take some time for the camera to be inserted, don't panic!
- You will need to make a fresh gain reference to be sure
Camera temperature not stable warning in SerialEM
- This is more a bug than a problem and mostly/only happens if you are running a script after a restart of the DM software and/or hardware.
- Just make a single view image with SerialEM (dark green panel, View) (Figure 1).
- Normally it will just have made the image and you can continue your previous operation.
- If it did not work, wait a few seconds and make again a new view image (Figure 1).
